

- Frontpage
- Image
- Authors
- Table of contents
- List of Abbreviations
- 1. Preface
- 2. Partners and collaborators
- 3. Workshop summary and outcome
- 3.1 Background of the workshop
- 3.2 Objectives of the workshop
- 3.3 Mapping of current knowledge for NAMs
- 3.3.1. Survey
- 3.3.2. Workshop poll
- 3.4. Summary of workshop
- 3.5. Conclusions
- 3.6. Specific recommendations from the workshop
- 3.7. Ongoing and follow-up NAMs activities
- 3.8. Workshop evaluation
- 4. Biographies of the speakers
- 5. Summary of presentations
- 5.1. Introduction to New Approaches Methodologies (NAMs)
- 5.2. Introduction to omics
- 5.3. Towards regulatory applications of molecular mechanistic data
- 5.4. An introduction to EPA CompTox Dashboard and GenRA Read-Across module
- 5.5. Challenges of NAMs in the real world 1
- 5.6. Challenges of NAMs in the real world 2
- 5.7. Plenary discussion training needs
- 5.8. Experience of ECHA in applying NAMs in a regulatory context
- 5.9. Omics-based grouping/read-across (G/RAx): A case study with azo dyes
- 5.10. Experience ECHA in applying NAMs in a regulatory context
- 5.11. Variability in repeat dose toxicity studies
- 5.10. Experience ECHA in applying NAMs in a regulatory context
- 5.12. Plenary discussion
- 5.13. Grouping and read-across case studies from EU-ToxRisk
- 5.14. ECHA’s experiences with NAMs in the assessment of chemical groups
- 5.15. EU-ToxRisk Case study 11
- 5.16. Orientation on relevant projects
- 5.17. H2020 PrecisionTox omics-based G/RAx case study in the making
- 6. Acknowledgements
- 7. Conflict of interest
- Appendix 1: Programme
- Appendix 2: Supplementary figures
- About this publication
MENU
Nordic Workshop on New Approach Methodologies (NAMs)
for Grouping and Read-Across under REACH and CLP
November 9th – 11th 2021
Dr Marcin W. Wojewodzic and Dr Monica Andreassen
Table of contents
This publication is also available online in a web-accessible version at https://pub.norden.org/temanord2022-526.
List of Abbreviations
ADME - absorption, distribution, metabolism, and excretion
AOP - Adverse Outcome Pathway
ASPIS cluster - “Animal-free Safety assessment of chemicals: Project cluster for Implementation of novel Strategies”. ONTOX, PrecisionTox and RISKHUNT3R are part of the ASPIS cluster
CLP - Classification, Labelling and Packaging
CSS - The Chemical Strategy for Sustainability
DNEL - Derived No-Effect Level
EC - European Commission
ECHA - European Chemicals Agency
G/RAx - Grouping/Read-Across
GMT - Group Management Team
IATA - Integrated Approaches to Testing and Assessment
KE – Key Event
LOAEL - Lowest-Observed-Adverse-Effect Level
MAD - Mutual Acceptance of Data
MIE – Molecular Initiating Event
MoA - Mode of Action
NAM - New Approach Methodology
NGRA - Next Generation Risk Assessment
NOAEL - No-Observed-Adverse-Effect Level
OECD - The Organisation for Economic Co-operation and Development
Omics - Transcriptomics, metabolomics, proteomics, epigenomics etc.
ONTOX - NAM Project funded by H2020 Programme. Coordinator is Prof Mathieu Vinken
PBPK - Physiological Based Pharmacokinetic modelling and simulation
POD - Point of Departure
PrecisionTox - NAM project funded by H2020 Programme. Coordinator is Prof John K. Colbourne
QSAR - Quantitative Structure-Activity Relationship
qIVIVE - Quantitative In Vitro to In Vivo Extrapolation
REACH - Registration, Evaluation, Authorisation and Restriction of Chemicals
RISKHUNT3R - NAM Project funded by H2020 programme. Coordinator is Prof Bob van de Water
1. Preface
New Approach Methodologies (NAMs) are a rapidly growing field of toxicology, offering a step away from the intensive use of animals in determining the hazard and toxicity of chemicals, and performing risk assessment for human health in the 21st century. Several major legal instruments have recently been put in place to reduce the use of animals in testing (including data sharing, grouping of chemicals, joint submission, and other adaptation possibilities of REACH, such as Annex XI). These initiatives have allowed NAMs to gain momentum, further helped by a growing interest in the implementation of Next Generation Risk Assessment (NGRA) principles.
There is an increasing trend to make regulations around NAMs and activities which will help ensure the smooth passage of NAMs into common practice. There have been government initiatives among regulatory agencies from North America, Europe, and Australia whose aim is to accelerate the pace of chemical risk assessment by using NAMs.
However, in the transition phase towards the use of these methods by legal authorities, the transfer of knowledge and competence within authorities is required. NAMs will be a very important point on the agenda in the EU for many years to come and, therefore, the Nordic countries need to be prepared for this shift. This transfer of knowledge should happen in parallel to the development of innovative and robust NAMs.
The Nordic workshop on NAMs was organised by the Norwegian Environment Agency, Nordic risk assessment project (NORAP) and the Nordic Classification Group (NKGI) and took place between the 9th and the 11th of November 2021. The workshop was organised as a webinar. The project was funded by the Nordic Council of Ministers and supported by the Nordic Working Group for Chemicals, Environment, and Health.
The target group of the workshop was hazard and risk assessors in the Nordic countries working with REACH and CLP. The aim of the workshop was to decrease the knowledge gap in the participants, to increase understanding of new methods utilising molecular mechanistic data (especially omics approaches), and to support their use in grouping and read-across methods. It also aimed to identify the needs in the Nordic countries for further capacity building, and guidance in NAM approaches, especially in a regulatory context.
Finally, the workshop aimed to highlight the current main regulatory challenges for NAMs, to exchange opinions and discuss possible ways forward for using NAMs, and to increase the level of knowledge and competence within the relevant authorities in the Nordic countries.
2. Partners and collaborators
The steering group of the workshop constituted Dr Hubert Dirven, Dr Birgitte Lindeman (both Norwegian Institute of Public Health, Norway), Dr Tomasz Sobański (European Chemicals Agency, Finland), Prof Mark R. Viant, Prof John K. Colbourne (both Michabo Health Science Ltd and the University of Birmingham, UK), Dr Daniel Borg (Swedish Chemicals Agency, Sweden and Nordic Risk Assessment Project), Tor Øystein Fotland (Norwegian Environment Agency, Norway and Nordic Risk Assessment Project) and Ann Kristin Larsen (Norwegian Environment Agency, Norway and Nordic Classification Group). The project manager and executive chair of the meeting was Marianne van der Hagen (Norwegian Environment Agency, Norway). The workshop was funded by the Nordic Council of Ministers.
Dr Marcin W. Wojewodzic and Dr Monica Andreassen (both Norwegian Institute of Public Health, Norway) were rapporteurs.
3. Workshop summary and outcome
3.1 Background of the workshop
The field of toxicology and toxicity testing of chemicals is rapidly advancing with the introduction of novel in chemico, in vitro, and in silico methods that have the potential to replace or complement the current animal experiment-based methods. Alongside the growth of New Approach Methodologies (NAMs) there is a growing demand for chemical hazard and risk assessors who have expertise in utilising the data generated from these methods. The participants represented various regulatory bodies of Finland, Norway, Denmark, and Sweden. Many of the participants perform risk assessments for their national authorities.
3.2 Objectives of the workshop
The workshop’s main aim was to increase the risk assessors’ understanding of how to use data produced by NAMs for assessing chemical hazards, within the Nordic countries. It also aimed to improve the assessors’ ability to use NAMs data in drafting proposals for regulatory actions under the chemical regulations REACH and CLP.
The European Chemicals Agency (ECHA) defines NAMs as methods that bring greater robustness, throughput and/or mechanistic knowledge into risk assessment, enabling more relevant decision making for human health and the environment. Aligned to this definition, the focus in the workshop was on new methods utilising molecular mechanistic data to support grouping and the read-across of chemicals.
This workshop also served to facilitate a discussion about the identified obstacles in the implementation of NAMs in regulatory processes and to identify strategies for increasing their use in read-across and grouping processes. The Norwegian Institute of Public Health (NIPH) drafted this report after the workshop, detailing the recommendations and needs for further actions by ECHA.
3.3 Mapping of current knowledge for NAMs
3.3.1. Survey
To identify the opportunities and challenges of NAMs in the real world, the participants were invited to complete an online survey upon registration for the workshop (5.5). The survey revealed that 64% of the respondents had some familiarity with the use of NAMs in a regulatory context. Respondents were familiar with grouping and read-across in regulatory decision making, and QSAR was the NAM most often used. Respondents identified both opportunities and challenges to include NAMs in grouping and read-across. Among opportunities seen by participants, NAM data could substitute regulatory toxicity observations of apical end points with observations of chemical modes-of-actions (MoAs) data, help improve hazard assessment, reduce animal testing, save money, and generate useful data when building a weight-of-evidence model. On the other hand, missing data as well as a lack of regulatory acceptance and validation of methods were reported as challenges in using NAMs for grouping and read-across
3.3.2. Workshop poll
To assess the participants’ prior knowledge about NAMs a comprehensive poll was conducted on day 1 of the workshop. The poll was repeated at the end of the workshop. The interpretation of the results is affected by unequal numbers of pre- and post- workshop responders (max of 36 versus 26 respectively). Due to this difference we cannot directly compare the pre- and post-workshop poll results.
Prior to the workshop, the participants were already familiar with some of the NAMs. As expected, many participants already had some experience with the ‘QSARs’, and ‘read-across’ techniques. Methods such as ‘IATAs‘ and ‘defined approach’ were also somewhat familiar to the participants. Most of the participants had already heard about the use of systematic reviews, but fewer had experience using it. Some participants also had experience with Adverse Outcome Pathways (AOPs) (Appendix 2, Figure 1).
The participants recognised opportunities of NAMs for many areas within toxicology in the next 5 years (Appendix 2, Figure 2). This was especially true for ‘read-across’ and ‘grouping’ methods that were of the main focus for this workshop. Also ‘sensitisation’, ‘irritation’, ‘endocrine disruptors’ and ‘genotoxicity’ were seen with large opportunities. After the workshop, the participants reported even larger opportunities in these areas (Appendix 2, Figure 5).
The participants were asked to identify the biggest hurdles in using NAMs for hazard assessments. They perceived the lack of formal regulatory requirements of NAMs and lack of validation of NAMs as big hurdles both before (Appendix 2, Figure 3) and after (Appendix 2, Figure 6) the workshop. This was not surprising, as these hurdles were discussed during the workshop. Lack of training in NAMs was also seen as a big hurdle suggesting a desire for more training and education. Further, the lack of industries submitting their data for reuse by others and site-by-site comparisons of NAMs and in vivo studies were indicated as hurdles. The polls also indicated that the participants had gained knowledge on how to retrieve NAM related data from international data series after the workshop (reported as less of a hurdle in the post-workshop poll). Knowledge about freely available databases was low among the participants in the pre-workshop poll (22%) (Appendix 2, Figure 4) but had increased after the workshop (73%) (Appendix 2, Figure 7). The participants were aware of various data sources like QSARs, ToxCast, EPA Dashboard, CompTox, AOP Wiki, Tox21, Vega and GenRA. Dashboard and GenRA stood out in particular after the workshop.
3.4. Summary of workshop
Introduction NAM definition is still evolving
As a newly emerging field, the definition of NAMs is still evolving. Three complementary definitions were presented by Prof Colbourne (5.1). He pointed out that these definitions are still under development, owing to scientific advances as well as the way different regulatory bodies perceive NAMs today. The definition in a regulatory context was more concrete and precise, probably recapitulating the legal needs behind future implementations of such methods. The scientific definition was much broader, reflecting the rapid pace of discoveries and innovation happening right now in this field. Independent of the presenters, the vision remained the same and advocates for the Replacement, Reduction, and Refinement of animal testing (the 3Rs) approach while maintaining the ultimate goal of protecting humans and the environment from chemical hazards. Dr Escher and Prof de Water also presented a common vision for human Next Generation Risk Assessment (NGRA), where hazard characterisation identification (currently done using animal tests) could be replaced by batteries of NAMs (5.13, 5.15).
NAM gains momentum
In the introductory lecture to NAMs, Prof Colbourne presented the main driving forces, in Europe and worldwide, for introducing NAMs into regulatory contexts. The main legal instruments to avoid animal testing are already in place, as reviewed by Prof Colbourne (5.1). He also highlighted the ongoing paradigm of moving away from extensive animal testing. This paradigm shift was reflected later on in the presentation of ongoing relevant projects in Europe (5.16). Most notably, the ASPIS cluster (Animal-free Safety assessment of chemicals: Project cluster for Implementation of novel Strategies) was created as a collaboration of the H2020 funded projects: ONTOX, PrecisionTox, RISK-HUNT3R. It represents Europe’s largest effort towards ‘the sustainable, animal-free and reliable chemical risk assessment of tomorrow’. These projects contain multiple elements of NAMs and, more importantly, they connect directly with regulatory bodies (5.9, 5.15, 5.16, 5.17). NAMs were clearly on ECHA’s radar as reviewed by Dr Sobański (5.8, 5.14) and in EPA, USA presented by Dr Paul Friedman (5.11). However, the motivation for these interests were slightly different. For ECHA it was the notion that REACH [1]Regulation (EC) No 1907/2006 of the European Parliament and of the Council on the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH) (Regulation (EC) No 1907/2006) and CLP [2]Regulation (EC) No 1272/2008 on the classification, labelling and packaging of substances and mixtures (CLP Regulation). (Regulation (EC) No 1272/2008) legislation were slightly disconnected from the operational level, while the EPA was more interested in using computational models to predict toxicity. Both groups are interested in speeding up the processes for hazard risk assessment and the prioritisation of chemicals.
Databases and data resources
This workshop was an arena for presenting databases with existing toxicological data that could be used for NAMs. In particular, the CompTox Chemicals Dashboard (comptox.epa.gov/dashboard) and the GenRA module were introduced (5.4). The data in the Dashboard is associated with more than 906k chemicals (as of February 2022) and Dr Paul Friedman demonstrated the practical uses of such databases in EPA research (5.11).
Use of omics in NAMs
In this workshop, omics approaches have been shown to be a powerful tool for grouping, read-across, and for obtaining a mechanistic understanding of a chemical’s MoAs. Prof Viant pointed out that while targeted approaches can be effective, non-targeted approaches can deliver new unknown biomarkers, and argued that both, or either, approaches should be used depending on the regulatory question being asked (5.2).
Multi-omics approaches have also been shown as important in comprehending the holistic picture of the MoAs of chemicals (5.2). These could bring a better understanding of key initiating events (5.2). Further examples were shown by Dr Escher (5.13) and Prof van Water (5.15) in human cell models using transcriptomics. The use of multi-omics was demonstrated by Prof Viant (5.9), where omics-based grouping and read-across was used in a case study with azo dyes.
During the workshop, ECHA presented how omics data are used in their dossiers. Dr Bouhifd (5.10) argued there is still very limited submission of omics data in general. This might be due to the lack of guidance or the reluctance by risk assessors and managers to use this type of information. He stated that the gaps in transparency within data analysis must be removed. More importantly, detailed descriptions of the pathways that link adversities need to be provided, otherwise the promise of mechanistic omics in toxicology risk assessment is not really fulfilled.
Another aspect to be considered is the relevance of the biological system to the endpoint of interest. While omics have great potential for measuring broad biological responses to a given stressor, this response needs to be measured on a model which is known to cover the toxicological space of interest, or there is a risk of it producing misleading data. The relevance of the model to a given stressor needs to be shown before omics approaches are used.
Footnotes
Case studies for NAMs
Several case studies were presented for using NAMs in this workshop. Prof Viant has demonstrated how the omics-based grouping and read-across, using a case study with azo dyes, could be instrumental in predicting MoAs, and could even connect these to apical end points (5.9). Here, mechanistic grouping using multi-omics was a robust method for grouping. Dr Escher presented case studies to demonstrate the integration process of NAMs for a human risk assessment for ‘repeated dose toxicity’ and ‘reproductive toxicity’, within a read-across assessment (5.13). As she demonstrated, based on transcriptomic profiles, the biological similarities of the analogues could be calculated based on the similarity of the gene expression patterns (5.13). Similar ideas were also presented by Prof van de Water (5.15).
State-of-the-art for using NAMs in regulation
During the workshop, NAMs in a regulatory perspective were thoroughly presented and discussed (5.8). Examples of the use of NAMs in regulatory context were given (5.8). Dr Sobański presented the motivation of ECHA to enter the world of NAMs, and their experience in applying NAMs in a regulatory context (5.8). He presented the future vision of ECHA for speeding up regulatory work using these approaches (5.8). The hurdles and gains from regulatory perspectives were thoroughly discussed. ECHA supports several activities via the APCRA initiative and contributes to multiple EU research programmes (e.g. EU-ToxRisk, ASPIS cluster, PARC) (5.8, 5.16).
Challenges in regulatory NAMs
The workshop clearly defined the gap between regulatory practices and state-of-the art approaches for NAMs. Many challenges in the hazard characterisation and assessment in regulatory contexts were identified during the workshop (5.8). Dr Sobański mentioned that although there are provisions in REACH for promoting the development of alternative methods for assessing the hazards of substances there are also significant hurdles to overcome (5.8). In particular, the information requirements in REACH (as well as classification criteria) refer to animal tests, and often to specific OECD in vivo test guidelines, indicated in the REACH Annexes. Regulatory use of NAMs would require the development of Integrated Approaches to Testing and Assessment (IATA) and further extrapolation to human safety assessment.
Needs for training in NAMs for regulators
Throughout the workshop, the regulatory need for training was demonstrated several times. This was shown in prior surveys and polls at different stages of the workshop (3.3, 3.4, 3.8, 5.5, 5.6, and 5.7).
3.5. Conclusions
- This workshop has strengthened the Nordic countries’ knowledge in the use of new methodologies in a regulatory context.
- The workshop raised awareness of the current knowledge gaps among the participants, and identified future training needs to be addressed by ECHA.
- It clearly identified the Nordic needs for capacity building and guidance to be provided by ECHA in the NAM field (especially in read-across and grouping, CLH-dossiers, case studies, and omics).
- The workshop contributed towards increasing the competence among authorities on how to apply NAMs in their regulatory work.
- It emphasised that these methodologies (used to support grouping and read-across approaches in particular) have the potential to replace, reduce and complement the use of traditional animal experiments, and that training is needed now to take full advantage of this.
- Participants were shown how NAMs could be used to support grouping and read-across approaches to reduce animal use, improve hazard assessment, support the evaluation of chemical substitution and deliver hazard information of data-poor substances.
- The workshop improved the Nordic authorities’ capacity to use results from NAMs for drafting new chemical regulations (contributing to the United Nations sustainable development goals relating to good health and clean environment).
- Finally, the workshop identified the need for guidance and additional competence building in how to apply NAMs data.
3.6. Specific recommendations from the workshop
- The identification of a key event may be useful for determining a chemical’s MoAs, but it is recommended that multiple key events tied to the same AOP are identified where possible as this would give far stronger evidence to predict an adverse outcome.
- It is recommended that key events observed at the molecular level should be at the centre of new safety regulations. Thus, generating a shift in focus away from the apical outcomes in experimental animals and towards perturbations of important pathways that lead to toxicity.
- Although molecular biomarkers and targeted gene panels are already part of some current regulatory paradigms, it is recommended that regulators move away from using single molecular biomarkers and towards large panels of biomarkers to predict chemicals’ MoAs.
- The use of untargeted omics is recommended for generating new knowledge about toxicity MoAs as omics can be used in the building of mechanistic models, including AOPs.
- It is also recommended that omics data are incorporated into conventional grouping approaches to compensate for any inconsistencies in the grouping hypothesis.
- A sole reliance on chemical structure and/or physical-chemical properties is not sufficient for robustly grouping chemicals. It is therefore recommended that a variety of molecular data types, including metabolomics and transcriptomics, are to be used for grouping.
- It is recommended that omics are used in bridging the gap between animal models and humans via similar MoAs.
- Further contributions to case studies are recommended, especially in areas where regulatory bodies meet with the scientific communities.
- A need to develop a predictive toxicology roadmap in REACH and CLP that describes a strategy to incorporate NAMs in the regulatory framework was identified.
- It is recommended that advice is provided in the guidance documents for REACH and CLP relating to omics and NAMs to ensure that they cover the topics most relevant to regulators.
- It is recommended that guidelines are provided for analysing and reporting omics data to ensure reproducibility and transparency between different labs for grouping. Any steps in the data analysis that lack transparency must be removed.
- It is recommended that workflows are established to enable molecular mechanistic data to be used for grouping approaches and prioritisation.
- The routine use of biologically based grouping alongside structure-based methods is also recommended.
- The submission of omics data by REACH registrants and assessors should be encouraged, and one way of achieving this would be by providing assessors with more guidance on this topic.
- Further research into the construction of detailed descriptions of molecular pathways that link to bioactivity and adversities is recommended.
- It is recommended that further training is provided in the use of NAMs with particular focus on groups of experts working together to encourage knowledge sharing and increase confidence.
3.7. Ongoing and follow-up NAMs activities
- From the survey and the workshop polls it is evident that more training and hands on examples are warranted (3.3, 3.4, 3.8, 5.5, 5.6, and 5.7).
- The omics reporting framework project is under development by OECD and involves the development of guidance documents for consistent reporting of omics data from various sources (5.2, 5.3). The project aims to develop a framework for the standardisation of reporting of omics data generation and analysis, to ensure that all of the information required to understand, interpret and reproduce an omics experiment and its results are transparent and available (FAIR principle).
- Ongoing OECD activities on omics: oecd.org/chemicalsafety/testing/omics.htm (5.6).
- ECHA is involved in multiple ongoing NAMs related projects aiming to translate NAMs into regulatory applications (5.8, 5.14)
- The APCRA initiative and consortium case studies, contributing to multiple EU research programmes (e.g. EU-ToxRisk, ASPIS cluster, PARC, 5.16)
- Involvement in implementation of the EU Chemical Strategy for Sustainability (CSS), using ToxCast/Tox21 assays and QSAR predictions under Group Management Team (GMT), and its contribution to the development and adoption of Defined Approaches
- Contributions to OECD Expert groups working on Transcriptomics and Metabolomics Reporting Framework and QSAR assessment framework to revise principles and establish an assessment framework.
- Promotion of NAMs via training about NAMs for ECHA staff and committees, industry, and the scientific community.
- The EU-ToxRisk project aims to deliver read-across procedures and to establish the starting point for hazard and risk assessment strategies for chemicals of scarce information (5.15)
- Multiple projects aim to use and improve the AOP framework as a departure point for NAMs (5.16).
- Michabo Health Science Ltd invited workshop participants (i.e. regulatory bodies) to collaborate on the grouping and read-across case studies (5.17).
3.8. Workshop evaluation
At the end of the workshop an online evaluation was held to collect the views of the participants on the overall quality of the workshop, the extent to which the workshop aims were met, the extent to which the information provided in the workshop will be useful in their future work, etc.
Overall, the participants were very satisfied with the workshop (average rating 4.4/5, Figure 1A), and most participants considered the workshop to be useful in their future work (Figure 1B). The majority of the participants (90%) identified a need for future training and/or guidance in the use of NAMs (Figure 1C), which included training and case studies on biostatistics, bioinformatics, and omics technologies for the grouping and read-across of substances for regulatory purposes as well as more training in omics technologies in general.
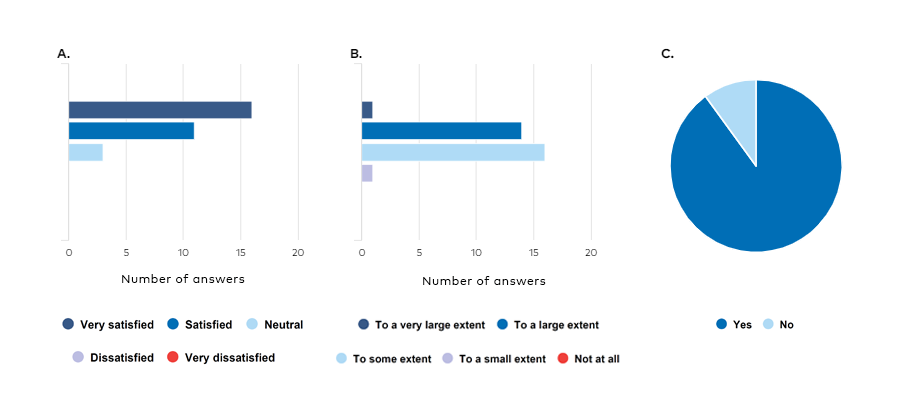
Figure 1. Results from the workshop evaluation A) How satisfied are you overall with the workshop? B) To what extent do you expect that the outcome of the workshop will be useful in your future work? C) Do you have any future needs for training and/or guidance in the use of NAMs
4. Biographies of the speakers
Dr Mounir Bouhifd is currently a regulatory officer at the European Chemicals Agency. He is member of the Alternative Methods Team of the computational Assessment unit. Part of his tasks is on the assessment of QSAR predictions. NAMs are another area of his work. Mounir was working on the development and application of Alternative methods and especially their validation, at the European Centre of Validation of Alternative methods (ECVAM). He was also a faculty member at the Johns Hopkins university.

Prof John K. Colbourne holds the inaugural Chair of Environmental Genomics at the University of Birmingham (UK). He is also an Adjunct Professor at the Mount Desert Island Biological Laboratory (USA), Guest Professor at Hebei University (China), co-founder and CSO of Michabo Health Science Limited, and co-founder of the international Environment Care Consortium (ECC), and of the Solve Pollution Network. This year, he incorporates the Environment Care Foundation (USA) as the registered charitable arm of the ECC. Previously, he served as genomics director of the Centre for Genomics and Bioinformatics at Indiana University (until 2012) receiving research funding from the U.S. NSF, NIH and DOE to help pioneer the application of genomics for the study of environmental health, primarily using the freshwater crustacean Daphnia - an evolutionary, ecological, and toxicological model system. This work resulted in Daphnia's designation as a biomedical model species by the U.S. National Institutes of Health and receiving the Royal Society Wolfson Research Merit Award. His research in Birmingham receives funding from NERC, BBSRC, Royal Society, FSA, U.S. NIEHS, and the European Commission that is focused on the application of genomics for environmental health protection by spearheading Precision Toxicology for obtaining comprehensive knowledge on the effects of synthetic chemicals and environmental pollutants on biology, using new and established genomic model species. He also serves on the UK Government’s Hazardous Substances Advisory Committee (HSAC), which provides expert advice on how to protect human health and the environment from potentially hazardous substances.

Dr Hubert Dirven is a leader of the Chemical Toxicology Unit, at the Norwegian Institute of Public Health, and he is involved in hazard and risk assessment for REACH chemicals and for chemicals in food. He is also involved in many of EU research project such as ONTOX, POLYRISK, EXIMIOUS and PARC. Hubert has previously worked as a toxicologist in the pharmaceutical industry.

Dr Sylvia E. Escher joined the Fraunhofer Institute of Toxicology and Experimental Medicine in Hanover in September 2006, where she currently leads the in silico toxicology department in the field of human risk assessment. Her interests include the development and maintenance of toxicological databases such as RepDose, and their use to develop and improve human risk assessment methods. Examples include the development of NAM supported read-across approaches and the TTC concept. Her team is currently developing AOPs for pulmonary fibrosis and a PBK model addressing in particular the integration of in vitro ADME properties of airborne compounds. She has published about >100 peer-reviewed articles, abstracts and book chapters.

Dr Katie Paul Friedman joined the Center for Computational Toxicology and Exposure in the Office of Research and Development at the US EPA in August 2016, where she is currently focused on application of NAMs to chemical safety assessment, with additional interests in uncertainty in alternative and traditional toxicity information, endocrine bioactivity and developmental neurotoxicity prediction, and in vitro kinetics. One of her roles in the Center is to run the ToxCast programmes. Previously, Dr Paul Friedman worked as a regulatory toxicologist at Bayer CropScience with specialties in neuro-, developmental and endocrine toxicity, and predictive toxicology. She has been actively involved in multi-stakeholder projects to develop AOPs, alternative testing approaches, and the regulatory acceptance of NAMs. Her laboratory background includes development of high-throughput screening assays, the combined use of myriad in vitro and in vivo approaches, including receptor-reporter and biochemical assays, primary hepatocyte cultures, and targeted animal testing paradigms, to investigate the human relevance of thyroid and metabolic AOPs using probe chemicals. Dr Paul Friedman received a Ph.D. in Toxicology from the University of North Carolina at Chapel Hill.
Dr Grace Patlewicz is currently a research chemist at the Center for Computational Toxicology & Exposure within the US EPA. She started her career at Unilever United Kingdom, before moving to the European Commission’s Joint Research Centre in Italy and then to DuPont in the United States. A chemist and toxicologist by training, her research interests have been focused on the development and application of QSARs and read-across for regulatory purposes. She has authored ~135 journal publications and book chapters, chaired various Industry groups and has contributed to the development of technical guidance for QSARs, chemical categories, and AOPs under various OECD work programmes.

Dr Magda Sachana is an Administrator within the Environment Health and Safety Division of the OECD’s Environmental Directorate since 2015. She manages the development and implementation of policies in the field of chemical safety and contributes to the OECD Test Guidelines, Pesticide and Hazard Assessment Programmes. Dr Sachana among other projects is coordinating the OECD project on omics reporting frameworks.

Dr Tomasz Sobański is a Team Leader for Alternative Methods Team at the European Chemicals Agency. He is working at ECHA for over 12 years focusing at development and application of the alternative methods in the regulatory processes. For many years he was a project manager of the OECD QSAR Toolbox while recent years he dedicated to NAMs and its applications in regulatory science. Tomasz co-authored over 30 publications and book chapters.

Mark R. Viant is Professor of Metabolomics at the University of Birmingham, UK, Executive Director of Phenome Centre Birmingham – a centre specialising in toxicometabolomics, and co-Founder/CEO of Michabo Health Science Ltd. He is also a past President of the International Metabolomics Society. His research focuses on developing and applying metabolomics in the field of human and environmental toxicology, with the goal to find novel molecular mechanistic solutions for industry and regulators in chemical safety science. He co-led the Ecetoc MEtabolomics standaRds Initiative in Toxicology (MERIT) project, and currently co-leads the omics activities within the OECD’s chemical safety programme and leads the Cefic MATCHING international ring-trial in toxicometabolomics. Mark has co-authored over 180 publications and his work has been recognised by the award of a 2015 Lifetime Honorary Fellowship of the International Metabolomics Society.

Bob van de Water is Professor of Drug Safety Sciences at the Leiden Academic Centre for Drug Research at Leiden University in the Netherlands. He has worked on molecular mechanisms of toxicity for over 30 years. His toxicogenomics research led to the discovery of biomarkers that have been integrated in fluorescent reporter test systems to qualify and quantify adverse cellular stress responses in relation to genotoxicity and severe cell injury. He was the coordinator of the Horizon2020 EU-ToxRisk project that focussed on mechanism-based testing strategies for read across. He currently coordinates the Horizon2020 RISK-HUNT3R project that will focus on NGRA strategies in the context of ab initio testing. Finally, he will be task leader in the Horizon Europe PARC programmes.

Dr Antony J. Williams joined the Center for Computational Toxicology and Exposure in the Office of Research and Development at the US EPA in May 2015. His interests include the aggregation and curation of chemical data, delivery of the center’s data to the scientific community, development of models to support physicochemical property prediction and development of software approaches to support non-targeted analysis. While his PhD is in Nuclear Magnetic Spectroscopy he moved into the field of cheminformatics and chemical information management over two decades ago. His focus has been on internet-based projects to deliver free-access community-based chemistry websites. He was one of the co-founders of ChemSpider that he started as a hobby project and is now hosted by the Royal Society of Chemistry. He is widely published with >300 peer-reviewed articles, book chapters and books.
5. Summary of presentations
5.1. Introduction to New Approaches Methodologies (NAMs)
By Prof John K. Colbourne
In his introductory talk, Professor John K. Colbourne presented various definitions of NAMs used by regulatory toxicologists. He has shown which technologies shaped the development of NAMs, and how legislative bodies played (and still play) a role in promoting the use of NAMs towards 21st century risk assessment. He conveyed how molecular toxicology can deliver mechanistic data, and how the molecular data can predict adversity. He described approaches used in comparative biology for the cross-species extrapolation of risk assessment results, advocating tiered approaches with both model organisms as well as cells.
New Approach Methodologies (NAMs) in toxicology
With the advent of NAMs, entering the portfolio of tools in regulatory processes, Prof Colbourne suggested that their effective uses in safety assessment may likely require a fundamental shift in our understanding of toxicology by focussing on MoAs. This will facilitate meaningful decisions around the hazards and risks of exposure to chemicals for human health and environment.
Definitions of New Approach Methodologies (NAMs)
The definition of NAMs is still developing and there were three working definitions provided at the workshop: I) ECHA regulators define NAMs as any approach for chemical hazard and risk assessment which can significantly contribute to throughput, robustness, and mechanistic knowledge, and which provide appropriate protection levels for human health and the environment. II) The UK Committee on Toxicology and Chemicals defines NAMs as high throughput screening omics and in silico computer modelling strategies. This also includes machine learning (with Artificial Intelligence) for the evaluation of hazard exposure. It advocates for the Replacement, Reduction, and Refinement of animal testing (the 3 Rs) approach. III) The USA EPA more broadly defines NAMs as any technology, methodology or approach, or combination that provides information on chemical hazard or risk assessment to avoid the use of animal testing, for instance omics derived NAMs. US priorities have also been set to reduce animal testing and to practically eliminate it by the year 2035.
The main legal instruments to avoid animal testing today
Today the testing proposals and third-party consultations require animal testing and reporting according to REACH (Regulation (EC) No 1907/2006) in EU. There exist five adaptations (NAMs) that have been used by industry when registering their compounds in their dossier for reporting under REACH. These include 1) the use of existing data, 2) the use of weight-of-evidence approaches, 3) information generated through quantitative structure activity relationships (QSARs), 4) in vitro test methods and 5) groupings of substances and read-across methods. Prof Colbourne showed that the grouping of substances and read-across methods are the most predominant form of NAMs used in the registration of chemicals today.
Paradigm of moving away from animal testing
The modern paradigm of moving away from animal testing was conceived in 2010, with a ground-breaking report that created the modern approach to performing hazard assessment [1]Gibb, S., Toxicity testing in the 21st century: a vision and a strategy. Reprod Toxicol, 2008. 25(1): p. 136–8.. It was commissioned in the United States by the National Academy of Sciences, Engineering and Medicine to address the backlog of chemical safety testing. It proposed an increase in throughput and the reduction of costs by utilising a mechanistic understanding of chemicals’ MoA.
This approach is designed specifically to assess the risk to humans posed by exposure to chemicals under the increased social demand to reduce or eliminate the use of animals (mammalian species in particular) for chemical safety testing. This is motivated by a desire to move away from making direct observations of adverse health effects (including the deaths of the animals used for toxicity testing), towards new, less harmful methodologies. These new methods are now enshrined in law and will likely continue to expand in the same way that animal testing laws have done for cosmetics.
The fundamental criteria that have been set by this report include, developing a more robust scientific basis for assessing the health effects of chemicals, providing a broad coverage of chemicals, chemical mixtures, outcomes, and life stages, reducing the cost and time of testing, and basing the decisions on human rather than rodent biology with more focus on relevant dose levels.
Prof Colbourne questions whether toxicity at environmentally relevant doses can truly be inferred from toxicity testing results at high doses and whether the toxicity observed in a mouse is truly predictive of the toxicity in humans. The proposed alternative solution to predict toxicity in humans is to measure the process of how exposure to chemicals perturbs normal biological functions, and to test a broad range of doses that set toxicity thresholds as points of departure (PODs) along the cellular response pathways that may cause adverse outcomes. This recommends that regulation refocuses hazard assessments on toxicity processes and MoAs expected to include early molecular changes, instead of on prescribed apical endpoints (such as animal deaths or reproduction outcome).
Toxicity as a biomolecular process
Prof Colbourne stressed that by recognising toxicity as a biomolecular process, regulatory toxicology can focus on understanding the potential harmful effects of chemicals at the level of the toxicity relevant pathway, which is defined here as a molecular response that, when sufficiently perturbed, is expected to result in an adverse health effect.
Proposals of the report
When this report was commissioned [2]Gibb, S., Toxicity testing in the 21st century: a vision and a strategy. Reprod Toxicol, 2008. 25(1): p. 136–8., there were different options for modern toxicity testing strategies that had been investigated by the National Academies. These were presented in detail by Prof Colbourne.
The report concluded that a combined tiered approach to toxicity testing (in vitro, in vivo, and in silico) may be the best option to modernise testing strategies in the near future. A combined approach, using both cell culture and whole organism testing would meet many criteria for new approaches discussed in the report. Mammalian systems are used only when compounds are likely to trigger pathways of unknown MoAs or have been flagged as potentially hazardous by toxicity pathway screening.
The proposed approach is intended to diversify the applicability of toxicity testing data, and to deliver a weight-of-evidence approach to risk assessment. This includes the systemic development of a tiered decision tree, selection of data, and evidence of toxicity from a limited suite of animal tests.
Weight-of-evidence refers to an approach that 1) Characterises chemicals based on their structure and function 2) Provides additional evidence of toxicity relating to hazards, and 3) Provides more data for dose response and extrapolation modelling. At each of these steps, the population base and human exposure data may be considered. In the end, the chemical regulator decides what data are needed for decision making but, at its centre, it is based on the knowledge of a chemical’s pathway.
The report proposes that tier one be exclusively focused on the evaluation of perturbations of pathways leading to toxicity rather than apical endpoints and may be done almost entirely in vitro. It also emphasises use of high-throughput approaches or cell lines of human origin, and to use medium throughput in vivo assays.
Paradigm shift with Tox21 programmes
This report led to a ground-breaking science programme in the United States called Tox21, which expanded on the report by recognising that chemical exposure experiments should be conducted on a broader range of animal species both at the cellular and organismal levels. The rationale for this is that chemical perturbations of the critical pathways are likely fundamental to animal biology including humans. The use of biomedical model species, such as zebrafish, fruit fly, or nematode worms, can improve throughput due to their small size and rapid reproduction rate. Furthermore, these species are regarded as non-sentient, and therefore not considered animals in a legal context. Therefore, greater throughput and the reduction of animal testing is achieved and toxicity testing on in vivo systems is used only to compensate for the drawbacks of in vitro research.
Transcriptomics and metabolomics
Prof Colbourne proposed that transcriptomics (quantitative measures of gene expression) and metabolomics (quantitative measures of metabolites) have the potential to reveal the chemical MoAs. With a large enough data set these leads to computational approaches that will either directly predict the effects of a chemical on human biology or prioritise the tests that need to be done on animals.
The use of in vitro testing is limited for detecting systemic toxicity. The physical behaviours of large multicellular networks and the interactions between diverse cells, tissues and organ types are complex making it impossible to gain a more complete understanding of the toxicity of a chemical from in vitro tests alone. Some of these drawbacks of in vitro tests can be overcome by testing on a suite of model species.
Footnotes
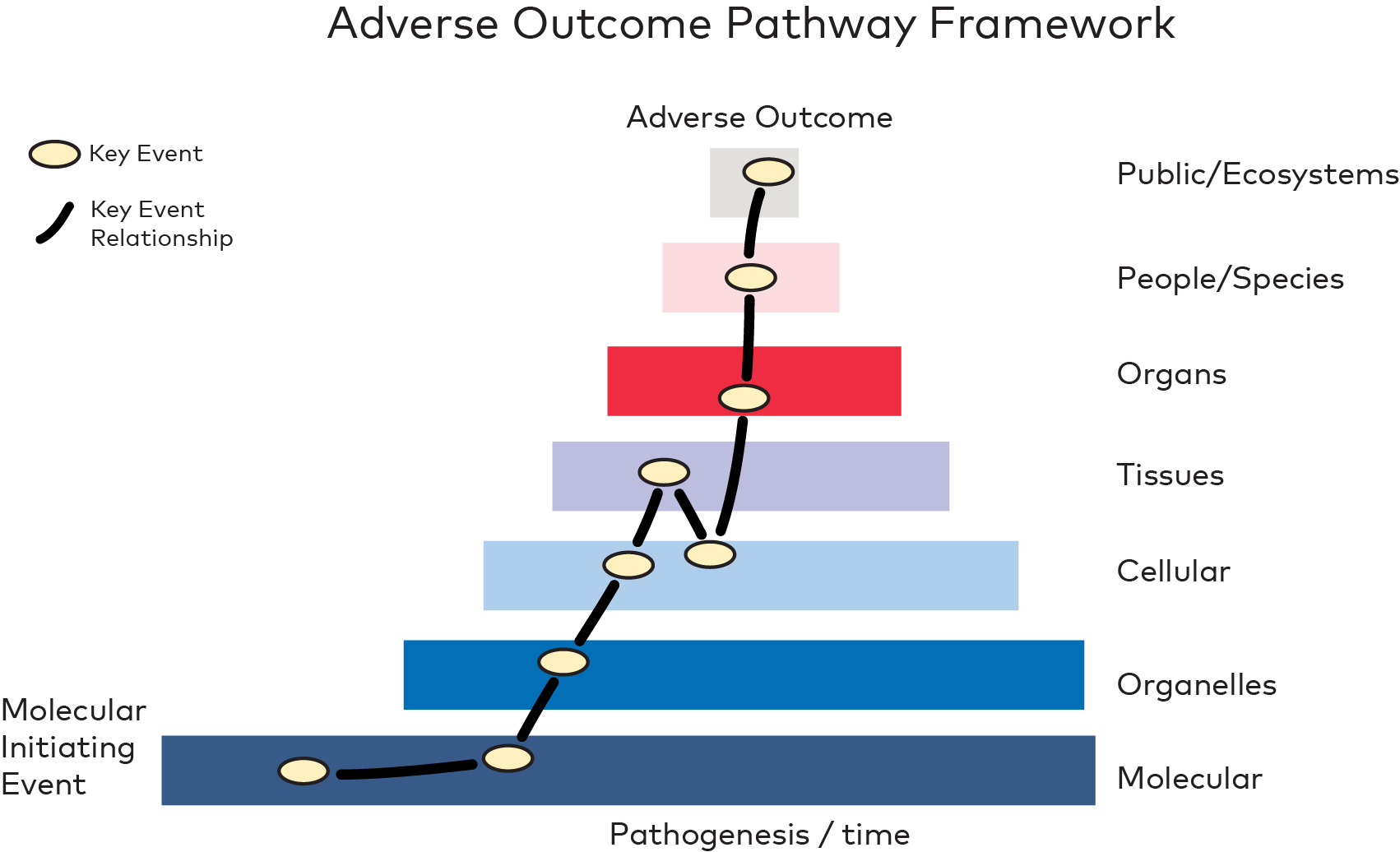
Figure 2. Adverse Outcome Pathways (AOPs) presented by John K. Colbourne after visualisation by Maurice Whelan (European Union Reference Laboratory for alternatives to animal testing (EURL ECVAM) at the JRC (Joint Research Centre of the European Commission), Italy, modified by Marcin W. Wojewodzic.
A mechanistic understanding of a specific toxicity pathway can be gained by analysing the key events that lead to AOPs for that chemical group (Figure 2). The utility of NAMs is expected to grow alongside the development of molecular key event databases, which connect the molecular signatures of toxicity to the associated chemical perturbations affecting all species, including humans.
The AOP is a flexible framework, centred around the key events. Prof Colbourne advocates that NAMs using early molecular key events in pathogenesis could be the future for risk assessment. Using this method, the observation of an organism's death or reproductive failure may no longer be necessary.
Key events are quantifiable as they are the fundamental changes in the biological condition of an organism. Key events set up the conditions leading to the occurrence of future downstream events, meaning that they, ultimately, are predictive of adverse outcomes. Furthermore, the observation of multiple key events associated with a given adverse outcome statistically improves the probability that that outcome will happen.
In the chain of key events leading to an adverse outcome, there is an initial earliest point (the molecular initiation event). This is the point at which biology intersects with chemistry. The molecular initiation event is a direct compound interaction with the biological molecules which ultimately leads to the adverse outcome. These sequences of key events are linked together across various levels of biological organisations through what are called key event relationships. The key event relationships are direct relationships that describe how one key event leads to another.
Finally, the adverse outcome is the furthest downstream key event in the chain. The adverse outcome is the key event that should determine the regulatory classification given to chemicals known to cause a corresponding molecular initiation event. Prof Colbourne argued that an improved understanding of key event chains will improve the ability of regulatory bodies to make decisions based on a better understanding of a given chemical’s MoA.
Prof Colbourne concluded by demonstrating an existing AOP that has been verified and accepted for skin sensitization. He reflected upon various areas of biological complexity at the molecular level, at the cellular level, at the organ level, and finally, at the whole organism level.
Conclusions
NAMs are a well-timed approach to toxicology because, ever since the implementation of REACH legislation, there's been a drive to step away from the intensive testing on animals to determine toxicity. There are some significant legal changes that have been recently put into place to reduce animal testing and accelerate NAMs development.
Identification of a key event may be useful for determining a chemical’s MoA, but the identification of multiple key events that are all tied to the same AOP gives far stronger evidence to predict an adverse outcome.
The regulatory context of 21st century toxicology is going to be a shift in focus away from the apical outcomes in experimental animals and towards these important perturbations of pathways leading to toxicity. The key events (most importantly, key events observed at the molecular level) and the development of risk assessment practices based on pathway perturbations will be at the centre of these changes. This will lead to the reinterpretation and rewriting of regulatory statutes under which risk assessments are conducted.
Questions, comments, and discussion
The public was interested to know how Prof Colbourne perceives the role of invertebrates in toxicology, when interpolating to human health. Prof Colbourne argued that by avoiding the use of mammals and fish for toxicological testing, we are pushed towards the use of invertebrates. In vitro methods can replace some in vivo testing, however there are still good reasons to observe the systemic perturbation of biological phenomena that are broadly shared by all animals. Prof Colbourne argued that research indicates that most of the pathways that are relevant to human health emerged very early in animal evolution. Therefore, he advocated invertebrates are viable substitutes to vertebrates for hazard assessment in terms of the identification of perturbed pathways. Many conservative pathways are shared with invertebrates as well.
5.2. Introduction to omics
By Prof Mark R. Viant
As an introduction to omics, Prof Viant focused his talk on molecular biomarkers, going from measuring single biomarkers to panels of thousands of biomarkers in a regulatory context. His presentation aimed to demystify the omics technology.
Single molecular biomarkers
OECD plays an important role in developing test guidelines and in the mutual acceptance of data. An accepted OECD guideline will be recognised by all 38 member states of the OECD. The OECD test guideline 408 repeated dose 90-day oral toxicity study in rodents was recently revised placing a greater emphasis on endocrine disruption. Prof Viant used endpoints in this revised guideline as a way of introducing molecular biomarkers, referring to the endogenous molecular metabolites thyroxine (T4) and triiodothyronine (T3). Their role in a mechanistic toxicity pathway is well understood, as they are responsive to thyroid pathway perturbation. Being included as a required endpoint in a standardised OECD test guideline, demonstrates that molecular biomarkers are in fact a part of the current regulatory paradigm.
Panels of molecular biomarkers
Another example presented by Prof Viant was an in vitro assay predicting skin sensitisation, the GARD®skin, which is under consideration for OECD test guideline programmes TGP 4.106. In contrast to assays built on single molecular biomarkers, this assay is based upon the readout of 200 genes. The GARD test methods make use of a machine learning algorithm (Support Vector Machine) to process genomic data. Such targeted assays that are measuring a molecular key event, whether it be metabolites or genes, are predicting a specific MoA. The next step is the concept of grouping assays together, to measure larger panels of biomarkers and be able to predict a broader range of MoAs. In 2018, US National Toxicology Program [1]Mav, D., et al., A hybrid gene selection approach to create the S1500+ targeted gene sets for use in high-throughput transcriptomics. PLoS One, 2018. 13(2): p. e0191105. published a gene biomarker panel with >1500 genes. Another example is the MTox700 metabolite biomarker panel predicting 722 human relevant metabolites associated with toxicity, adverse outcomes, and disease [2]Sostare, E., et al., Knowledge-driven approaches to create the MTox700+ metabolite panel for predicting toxicity. Toxicol Sci, 2022..
Demystifying omics
Omics techniques aim to characterise and quantify biological molecules that translate into knowledge about the functioning of an organism. Transcriptomic, metabolomic, proteomic, genomic, epigenomics are the most common omics techniques used today. Measuring a large panel of predefined molecular biomarkers that cover a broader range of perturbations, i.e. measuring characterised MoAs, is referred to as targeted omics. However, this approach will only measure what is already known today and may miss chemically induced biologically relevant effects that are yet unknown. To address this issue, the next level is therefore the application of untargeted omics, exploring both characterised and uncharacterised MoAs and providing broader views of the toxicological response to a chemical. Untargeted omics includes measuring all possible genes or metabolites, not just those that are associated with a particular MoA.
It is a general point of view that there is a unidirectional travel of information from the genome through the transcriptome. The word transcriptome refers to expressed genes (mRNA), the proteome refers to all proteins, and the metabolome refers to all metabolites. To study effects of chemicals on biological systems, Prof Viant emphasised the importance of using the powerful combination of expressed genes and metabolic biomarkers (multi-omics approach), to reduce uncertainty and increase confidence in the prediction of an adverse outcome.
From mechanistic data towards regulatory application
The apparent limitation of any assay that measures a single MoA is that many individual assays are needed to cover a wide range of MoAs in a hazard or risk assessment. Nevertheless, it is a good starting point. From the previous examples, we know that the use of molecular biomarkers and targeted gene panels are already in use, accepted or under consideration in regulatory paradigms. The use of untargeted omics is still at an early stage, at least in the context of regulatory toxicology, and there are challenges that need to be addressed, including improved mechanistic anchoring of the omics data.
Finally, Prof Viant highlighted that molecular biomarker assays are built on previous knowledge of biochemical pathways in toxicology. Gene assays like GARD®skin, however, originated from an untargeted omics study and the use of machine learning approaches to identify the 200 genes included in the assay. Thus, another role of omics will be to discover biomarkers in the first place.
Conclusions
Molecular biomarkers and targeted gene panels are already part of the current regulatory paradigm. Moving from single molecular biomarkers to panels of thousands of biomarkers, enables a broader prediction of MoAs. The application of untargeted omics, exploring uncharacterised MoAs and providing an even more comprehensive view of the toxicological response to a chemical, is promising but still at an early stage in the context of regulatory toxicology.
Footnotes
- ^ Mav, D., et al., A hybrid gene selection approach to create the S1500+ targeted gene sets for use in high-throughput transcriptomics. PLoS One, 2018. 13(2): p. e0191105.
- ^ Sostare, E., et al., Knowledge-driven approaches to create the MTox700+ metabolite panel for predicting toxicity. Toxicol Sci, 2022.
5.3. Towards regulatory applications of molecular mechanistic data
By Prof Mark Viant
The focus of Prof Viant’s talk was on grouping and read-across using molecular mechanistic data. He also gave a brief introduction to the determination of potency via benchmark dosing.
Concept for grouping using molecular mechanistic data
It has been pointed out that a sole reliance on chemical structure is not enough for grouping of chemicals. Prof Viant started his presentation introducing the concept for the grouping of chemicals using molecular mechanistic data. Firstly, the transcriptome is condition dependent, and chemicals acting via different MoAs induce different sets of gene biomarkers. This implicates that those chemicals acting via the same MoA should induce similar gene biomarkers. The concept can also be applied to metabolomes, which are also condition dependent, i.e., chemicals acting via different MoAs induce different metabolic biomarkers, suggesting that chemicals acting via the same MoA should induce similar metabolic biomarkers. However, the thought of using molecular mechanistic data in grouping is not new. The term bioprofiling, which includes transcriptomics and metabolomics, has already been used by OECD (and others) in existing guidance documents for grouping and read-across[1]OECD, Guidance on Grouping of Chemicals, Second Edition. 2017..
Deficiencies
The ECHA report on alternatives to testing on animals for the REACH Regulation[2]ECHA, The use of alternatives to testing on animals for the REACH Regulation. Fourth report (2020) under Article 117(3) of the REACH Regulation. 2020. addresses shortcomings in grouping and read-across. The most common shortcomings include the lack of, or low quality of, supporting data, and limitations in the hypothesis and justification of the toxicological prediction. The report specifically mentioned that to increase robustness and regulatory acceptance for human health endpoints, additional data is needed, particularly related to toxicological mechanism and ADME properties. Furthermore, it is suggested that NAMs (for example high-throughput in vitro screening) have the potential to further substantiate the hypothesis of read-across approaches. Prof Viant underlines the word ‘potential’ to remind everyone that there are still several issues to be resolved.
How to bring molecular mechanistic data into grouping and read across?
According to Prof Viant, bringing molecular mechanistic data into the procedure does not involve a fundamental change in the grouping and read-across paradigm. New data in terms of molecular biomarkers would be added to the multi-step process of grouping and read-across to build confidence in the formation of the group or category, while the rest of the paradigm stays unchanged. As an example, the process could involve the measurement of gene expression and metabolites to characterise biological response to chemical exposure, calculating the similarity of the gene expression and metabolites between each pair of chemicals, and visualising the similarities to form groups of chemicals. Several papers have been published on this approach in recent years [3]Nakagawa, S., et al., Grouping of chemicals based on the potential mechanisms of hepatotoxicity of naphthalene and structurally similar chemicals using in vitro testing for read-across and its validation. Regulatory Toxicology and Pharmacology, 2021. [4]Sperber, S., et al., Metabolomics as read-across tool: An example with 3-aminopropanol and 2-aminoethanol. Regulatory Toxicology and Pharmacology, 2019..
Determination of potency via benchmark dosing
Benchmark dose (BMD) modelling is used to estimate the point of departure to define a safe dose. Increasingly, this approach has been applied to molecular data, looking at the level of a particular molecular response as a function of dose. Prof Viant gave an example using a molecular biomarker associated with energy metabolism to illustrate the possibility to estimate potencies.
Conclusions
There is wide acceptance that a sole reliance on chemical structure and/or physical-chemical properties is insufficient for robustly grouping chemicals. Different types of molecular data, including metabolomics and transcriptomics, can be used to provide confidence to grouping and in principle enable a more reliable read-across. Metabolomics data has previously been submitted to ECHA to support read-across and data gap filling for 3-aminopropan-1-ol.
Footnotes
- ^ OECD, Guidance on Grouping of Chemicals, Second Edition. 2017.
- ^ ECHA, The use of alternatives to testing on animals for the REACH Regulation. Fourth report (2020) under Article 117(3) of the REACH Regulation. 2020.
- ^ Nakagawa, S., et al., Grouping of chemicals based on the potential mechanisms of hepatotoxicity of naphthalene and structurally similar chemicals using in vitro testing for read-across and its validation. Regulatory Toxicology and Pharmacology, 2021.
- ^ Sperber, S., et al., Metabolomics as read-across tool: An example with 3-aminopropanol and 2-aminoethanol. Regulatory Toxicology and Pharmacology, 2019.
Towards regulatory application of molecular mechanistic data – part II
By Prof John K. Colbourne
Prof Colbourne gave a talk about detecting chemicals MoA and how to extrapolate across species by embracing the genetic similarities and differences in an evolutionary context.
Genetic similarities and differences across species
Most of our traits are shared with other animal species, inherited from common ancestors driven by a long history of animal evolution. An important question is to what extent can we rely on genomic observations made in invertebrates to predict the health effects of chemical hazards in humans? If genes are in fact responsible for coding elements that are disrupted by chemicals and therefore leading to toxicity, toxicology can focus cross-species extrapolation of toxicity pathways based on the knowledge of the evolutionary conservation of genes and their functions (comparative biology of animal genomes). Genes that are most strongly linked to adverse outcomes are disproportionately shared among all animals including humans. Differences among species can either be qualitative or quantitative; Qualitative differences are attributed to gains or losses of traits, including pathways relevant for toxicity, while quantitative differences can be attributed to genetic variation, either segregating within or among population of the same species, or non-segregating genetic variation among species.
Chemical modes of action (MoAs) and extrapolation across species
Prof Colbourne presented three examples using existing data on how chemical MoAs is detected and extrapolated across species by embracing the genetic similarities and differences that we have in our evolutionary history.
In the first example it was demonstrated how qualitative differences, in terms of metabolic pathways diverge among species. The benzoyluera class of insecticides are designed by industry to inhibit chitin synthesis. Insects have a chitin exoskeleton whereas vertebrates do not, and this rational was used to design this class of insecticides. The toxicity of this insecticide should be dependent upon whether or not an animal species contains a chitin synthesis pathway in its genome. However, systematic review of literature on the sequencing of animal genomes including humans, other vertebrates and invertebrates revealed that the chitin synthase gene and the pathway originated by an ancestor to both vertebrates and invertebrates at the base of the animal phylogeny. However, by evolutionary chance, the chitin synthase gene was lost in the branch leading to humans and other mammals. Prof Coulbourne referred to the fact that these insecticides is not predicted to be toxic to humans as an evolutionary “flip of the coin”.
In the second example, Prof Colbourne demonstrated quantitative differences in terms of rate variation in metabolic pathways. The metabolism of certain compounds is dependent on the presence or the absence of the pathway itself, or by the varying efficacy of the enzymes that are encoded by variation found among animal genomes including the human genome.
The third example referred to how genetic variation could influence toxicity. Genetic variation within and among animal population (including humans) control processes that affects exposure, dose, toxicokinetics and toxicodynamics. For example, a simple amino acid substitution may radically alter the efficiency of an enzyme and hence influence toxicity. In a case study, it was demonstrated how genetic variation in human hepatocytes from eight individuals affected the capacity to metabolise and detoxify inorganic arsenic.
Based on these examples, Prof Colbourne emphasises that safety testing on animals, exclusively based on observed apical endpoints can be either protective of human health or misleading, depending on the test animal and its genetic basis for toxicity. NAM approaches may be helpful in terms of having a more nuanced understanding of the risks that chemicals pose to humans.
Finally, Prof Colbourne talked about prediction of liver toxicity and MoAs using metabolomics of in vitro HepG2 cells [1]Ramirez, T., et al., Prediction of liver toxicity and mode of action using metabolomics in vitro in HepG2 cells. Arch Toxicol, 2018. 92(2): p. 893–906.. Here, compounds were grouped according to their presumed molecular MoA.
Conclusions
In conclusion, Prof Colbourne advocates that omics can redirect testing to achieve the following:
- Improve certainty in selecting critical endpoints through building and supporting mechanistic information
- Enhance understanding of whether adverse effects observed in animals are likely to occur in humans via similar MoAs /molecular key events
- Guide in selecting appropriate risk assessment approaches (such as threshold/quantitative or non-threshold/qualitative approaches)
- Support chemical grouping and read-across
Footnotes
- ^ Ramirez, T., et al., Prediction of liver toxicity and mode of action using metabolomics in vitro in HepG2 cells. Arch Toxicol, 2018. 92(2): p. 893–906.
Guidance document for consistent reporting of omics data from various sources
By Prof Mark Viant
The omics reporting framework project is being conducted by OECD and involves the development of guidance documents for consistent reporting of omics data from various sources. The goal is to develop a framework for the standardisation of reporting of omics data generation and analysis, to ensure that all of the information required to understand, interpret and reproduce an omics experiment and its results are available. Prof Viant emphasised that the purpose is to ensure that sufficient information is available to enable an evaluation of the quality of the experimental data and interpretation, and support reproducibility, but not to stipulate the methods of data analysis or interpretation.
Covering the two omic approaches, the transcriptomics reporting framework (TRF) led by Joshua Harrill (USEPA) and Carole Yauk (formerly of Health Canada, now at the University of Ottawa), and the metabolomic reporting framework (MRF) led by Prof Viant (University of Birmingham, UK) are integrated in this work. Both the TRF and MRF are harmonised, providing a reporting template and a narrative guidance. In connection to that, Prof Viant promoted the paper by [1]Harrill, J.A., et al., Progress towards an OECD reporting framework for transcriptomics and metabolomics in regulatory toxicology. Regul Toxicol Pharmacol, 2021. 125: p. 105020. on the progress towards an OECD reporting framework for transcriptomics and metabolomics in regulatory toxicology.
To improve the guidance document, extensive trialling has been conducted via case studies with ‘data submitter’ and ‘end user’ teams, with comparison of the two sets of results in a concordance analysis. Moreover, trials are ongoing in a Cefic-funded project to demonstrate that multiple labs, each analysing and reporting omics data from a single toxicity study, can arrive at the same conclusion for grouping eight chemicals (cefic-lri.org/projects/c8-assessing-the-repeatability-of-metabolomics-within-a-regulatory-context-through-a-multi-laboratory-ring-trial/). This multiple lab ring-trial will also use the new MRF reporting framework.
Footnotes
- ^ Harrill, J.A., et al., Progress towards an OECD reporting framework for transcriptomics and metabolomics in regulatory toxicology. Regul Toxicol Pharmacol, 2021. 125: p. 105020.
5.4. An introduction to EPA CompTox Dashboard and GenRA Read-Across module
By Dr Antony J. Williams and Dr Grace Patlewicz
Vision of using CompTox for NAMs in computational toxicology
The vision of the computational toxicology research methods (NAMs) is to speed up prioritisation, and to decrease time and costs for chemical hazard evaluation. Dr Williams and Dr Patlewicz, introduced the publicly accessible CompTox Chemicals Dashboard (https://comptox.epa.gov/dashboard), followed by hands-on demonstrations in using this resource for different applications. The Dashboard provides public access to many of the databases developed by the Center for Computational Toxicology and Exposure at the US Environmental Protection Agency, as part of their Chemical Safety for Sustainability Research Program. It represents data generated and assembled over the last 20 years, initiated with the DSSTox database [1]Richard, A.M. and C.R. Williams, Distributed structure-searchable toxicity (DSSTox) public database network: a proposal. Mutat Res, 2002 499(1): p. 27–52., with an overarching goal to deliver a computational toxicology platform where the innovation of alternative ways of evaluating chemicals for health risks can flourish.
Overview of the CompTox
The data in the dashboard is associated with more than 906k chemicals (as of February 2022). Significant efforts are expended in manual curation of the data, as exemplified by the investment in curating and expanding the DSSTox chemistry data [2]Grulke, C.M., et al., EPA's DSSTox database: History of development of a curated chemistry resource supporting computational toxicology research. Comput Toxicol, 2019. 12.. However, multiple other databases integrate into the Dashboard including, but not limited to, the invitrodb database associated with ToxCast (epa.gov/chemical-research/exploring-toxcast-data-downloadable-data) and data associated with exposure predictions (epa.gov/chemical-research/rapid-chemical-exposure-and-dose-research). The dashboard contains information about individual assays from ToxCast/Tox21 studies with a rich interface, with the detail of individual data points for assays, including fitting procedures to the models, and updates to the modelling part as the science progresses. The examples of use models derived from the data of these can be found e.g. CERAPP [3]Mansouri, K., et al., CERAPP: Collaborative Estrogen Receptor Activity Prediction Project. Environ Health Perspect, 2016. 124(7): p. 1023–33. and CoMPARA [4]Mansouri, K., et al., CoMPARA: Collaborative Modeling Project for Androgen Receptor Activity. Environ Health Perspect, 2020. 128(2): p. 27002. projects. For more details about the ToxCast/Tox21 bioactivity data see lecture of K.P. Friedman, and “ToxCast Chemical Landscape: Paving the Road to the 21st Century Toxicology” [5]Richard, A.M., et al., ToxCast Chemical Landscape: Paving the Road to 21st Century Toxicology. Chem Res Toxicol, 2016. 29(8): p. 1225–51..
Chemical substances added into the underlying DSSTox database are identified based on programmes of interest to the agency and are harvested from regulatory documentation, public domain databases, literature articles and other resources. The manual curation process is to ensure quality mappings between substance identifiers and associated chemical structures has been described in detail [6]Grulke, C.M., et al., EPA's DSSTox database: History of development of a curated chemistry resource supporting computational toxicology research. Comput Toxicol, 2019. 12..
Introduced in April 2016, the Dashboard [7]Williams, A.J., et al., The CompTox Chemistry Dashboard: a community data resource for environmental chemistry. J Cheminform, 2017. 9(1): p.61. was developed with the intention of providing an intuitive, searchable platform that would provide detailed information about the chemical of interest, its properties and structure while also linking to source information for given parameters. Since its initial release multiple layers of functionality have been added with each incremental release. For example, the Executive Summary gives an overview of toxicity-related information such as quantitative values, physicochemical properties, links to known AOPs, or in vitro bioactivity summary plots (see Bisphenol A as an example comptox.epa.gov/dashboard/chemical/executive-summary/DTXSID7020182).
While there are 100s of thousands of experimental data points associated with human and ecological toxicity data (i.e. the ToxVal database), physicochemical properties and fate and transport data, when these data are not available then QSAR prediction models have been used to generate predicted data. These include Toxicity Estimation Software Tool (TEST) predictions (epa.gov/chemical-research/toxicity-estimation-software-tool-test), OPEn structure-activity/property Relationship App (OPERA) [8]Mansouri, K., et al., OPERA models for predicting physicochemical properties and environmental fate endpoints. J Cheminform, 2018. 10(1): p.10. [9]Mansouri, K., et al., Open-source QSAR models for pKa prediction using multiple machine learning approaches. J Cheminform, 2019. 11(1): p.60., as well as commercial models. These prediction models are updated overtime with additional data and new predictions are added into future releases.
The Dashboard also provides access to multiple forms of exposure data including the Chemical and Products Database (CPDat) [10]Dionisio, K.L., et al., The Chemical and Products Database, a resource for exposure-relevant data on chemicals in consumer products. Sci Data, 2018. 5: p. 180125., exposure modelled data based on the Systematic Empirical Evaluation of Models approach (SEEM) [11]Ring, C.L., et al., Consensus Modeling of Median Chemical Intake for the U.S. Population Based on Predictions of Exposure Pathways. Environ Sci Technol, 2019. 53(2): p. 719–732. and predicted functional use [12]Phillips, K.A., et al., High-throughput screening of chemicals as functional substitutes using structure-based classification models. Green Chem, 2017. 19(4): p. 1063–1074.. To support exposure monitoring using mass spectrometry (MS) approaches (including non-targeted analysis) the chemical structures have been processed into a form known as “MS-Ready” [13]McEachran, A.D., et al., "MS-Ready" structures for non-targeted high-resolution mass spectrometry screening studies. J Cheminform, 2018. 10(1): p. 45. and have been used as the basis of many MS studies in recent years [14]McEachran, A.D., et al., Revisiting Five Years of CASMI Contests with EPA Identification Tools. Metabolites, 2020. 10(6). [15]Ulrich, E.M., et al., EPA's non-targeted analysis collaborative trial (ENTACT): genesis, design, and initial findings. Anal Bioanal Chem, 2019. 411(4): p. 853–866..
The Abstract Sifter literature search module [16]Baker, N., T. Knudsen, and A. Williams, Abstract Sifter: a comprehensive front-end system to PubMed. F1000Res, 2017. 6. has also been integrated into the Dashboard as well as searching strategies using other internet sources have been also implemented. These take into account the complexity associated with the many potential synonyms and identifiers associated with chemicals and allows for fine tuning queries based on real-time retrieval of data from PubMed (based on CAS numbers and other identifiers).
There are many flexible searches possible through the dashboard including chemicals (based on names and identifiers such as CAS RN), products and use categories, and assay and gene searches associated with ToxCast/Tox21 assays.
Batch searching [17]Lowe, C.N. and A.J. Williams, Enabling High-Throughput Searches for Multiple Chemical Data Using the U.S.-EPA CompTox Chemicals Dashboard. J Chem Inf Model, 2021. 61(2): p. 565–570. is also supported which allows users to query for data associated with up to ten thousand chemicals (for the present version) and download the data into standard formats including CSV and Excel.
A recent example of how the dashboard can be applied to supporting risk assessment was recently published: “Sourcing data on chemical properties and hazard data from the US-EPA CompTox Chemicals Dashboard: A practical guide for human risk assessment” [18]Williams, A.J., et al., Sourcing data on chemical properties and hazard data from the US-EPA CompTox Chemicals Dashboard: A practical guide for human risk assessment. Environ Int, 2021. 154: p. 106566. .
Footnotes
- ^ Richard, A.M. and C.R. Williams, Distributed structure-searchable toxicity (DSSTox) public database network: a proposal. Mutat Res, 2002 499(1): p. 27–52.
- ^ Grulke, C.M., et al., EPA's DSSTox database: History of development of a curated chemistry resource supporting computational toxicology research. Comput Toxicol, 2019. 12.
- ^ Mansouri, K., et al., CERAPP: Collaborative Estrogen Receptor Activity Prediction Project. Environ Health Perspect, 2016. 124(7): p. 1023–33.
- ^ Mansouri, K., et al., CoMPARA: Collaborative Modeling Project for Androgen Receptor Activity. Environ Health Perspect, 2020. 128(2): p. 27002.
- ^ Richard, A.M., et al., ToxCast Chemical Landscape: Paving the Road to 21st Century Toxicology. Chem Res Toxicol, 2016. 29(8): p. 1225–51.
- ^ Grulke, C.M., et al., EPA's DSSTox database: History of development of a curated chemistry resource supporting computational toxicology research. Comput Toxicol, 2019. 12.
- ^ Williams, A.J., et al., The CompTox Chemistry Dashboard: a community data resource for environmental chemistry. J Cheminform, 2017. 9(1): p.61.
- ^ Mansouri, K., et al., OPERA models for predicting physicochemical properties and environmental fate endpoints. J Cheminform, 2018. 10(1): p.10.
- ^ Mansouri, K., et al., Open-source QSAR models for pKa prediction using multiple machine learning approaches. J Cheminform, 2019. 11(1): p.60.
- ^ Dionisio, K.L., et al., The Chemical and Products Database, a resource for exposure-relevant data on chemicals in consumer products. Sci Data, 2018. 5: p. 180125.
- ^ Ring, C.L., et al., Consensus Modeling of Median Chemical Intake for the U.S. Population Based on Predictions of Exposure Pathways. Environ Sci Technol, 2019. 53(2): p. 719–732.
- ^ Phillips, K.A., et al., High-throughput screening of chemicals as functional substitutes using structure-based classification models. Green Chem, 2017. 19(4): p. 1063–1074.
- ^ McEachran, A.D., et al., "MS-Ready" structures for non-targeted high-resolution mass spectrometry screening studies. J Cheminform, 2018. 10(1): p. 45.
- ^ McEachran, A.D., et al., Revisiting Five Years of CASMI Contests with EPA Identification Tools. Metabolites, 2020. 10(6).
- ^ Ulrich, E.M., et al., EPA's non-targeted analysis collaborative trial (ENTACT): genesis, design, and initial findings. Anal Bioanal Chem, 2019. 411(4): p. 853–866.
- ^ Baker, N., T. Knudsen, and A. Williams, Abstract Sifter: a comprehensive front-end system to PubMed. F1000Res, 2017. 6.
- ^ Lowe, C.N. and A.J. Williams, Enabling High-Throughput Searches for Multiple Chemical Data Using the U.S.-EPA CompTox Chemicals Dashboard. J Chem Inf Model, 2021. 61(2): p. 565–570.
- ^ Williams, A.J., et al., Sourcing data on chemical properties and hazard data from the US-EPA CompTox Chemicals Dashboard: A practical guide for human risk assessment. Environ Int, 2021. 154: p. 106566.
Generalised Read-Across (GenRA)
Dr Patlewicz presented the Generalised Read-across (GenRA) approach which predicts toxicity as a similarity-weighted activity of nearest neighbours based on chemistry and/or bioactivity descriptors [1]Shah, I., et al., Systematically evaluating read-across prediction and performance using a local validity approach characterized by chemical structure and bioactivity information. Regul Toxicol Pharmacol, 2016 : p. 12–24.. The goal of the first version of GenRA (GenRA v1.0) was to establish an objective performance baseline for read-across and quantify the uncertainties in the predictions made. The approach facilitates the inclusion of other NAMs data streams such as transcriptomics and phenotypic profiling Overall, GenRA aims to move towards an objective read-across approach where uncertainties and performance can be quantified.
The initial version of GenRA relied upon chemical descriptors to predict binary toxicity values but research is ongoing to characterise other contexts of similarity (e.g. mechanistic, reactivity, metabolism) and quantify their contribution in predicting in vivo toxicity outcomes. The most recent research has explored the use of targeted transcriptomics data [2]Tate, T., et al., Repeat-dose toxicity prediction with Generalized Read-Across (GenRA) using targeted transcriptomic data: A proof-of-concept case study. Computational Toxicology, 2021: p. 100171. and the use of metabolic predictions [3]Boyce, M., et al., Comparing the performance and coverage of selected in silico (liver) metabolism tools relative to reported studies in the literature to inform analogue selection in read-across: A case study. Computational Toxicology, 2022. : p. 100208 GenRA v2.0, released just after this workshop is a rebuild of v1.0 with updated data and was created as a standalone web application. It maintains a link to the Dashboard such that GenRA can be accessed from the Dashboard as well as from its own direct entry point (comptox.epa.gov/genra/). GenRA v3.0 was just released (February 2022) with additional enhancements including a structure drawing palette to introduce chemicals not necessarily registered within the DSSTox database as well as the ability to use custom fingerprints to search for analogues. A python package (genra-py) was also released in March 2021 to facilitate batch processing using user specific datasets (github.com/i-shah/genra-py). A sample case study that allows an end-user to test out the package in a browser without any installation is also available (https://github.com/patlewig/UNC_Rax). The case study walks through an example dataset for acute oral toxicity and showcases how the package can be used to recreate an analysis were originally published [4]Helman, G., et al., Generalized Read-Across (GenRA): A workflow implemented into the EPA CompTox Chemicals Dashboard. ALTEX, 2019 : p. 462–465
Questions, comments, and discussion
Seven people out of 40 had used the Dashboard before participating. Participants appreciated the presented content of the Dashboard and wondered if there will be a publication with guidelines for best practice using GenRA v2. Whilst a manuscript was published following the initial release of GenRA v1. [5]Helman, G., et al., Generalized Read-Across (GenRA): A workflow implemented into the EPA CompTox Chemicals Dashboard. ALTEX, 2019. p. 462–465. no publication with guidelines for using GenRA v2 is planned. A short user manual is available to guide the end user in navigating the application. If there is strong interest in such a publication, we would welcome feedback in terms of what participants would find most useful. Our current focus is to continue researching ways and means of how different similarity contexts can be characterised to refine predictions.
There were questions from participants about the detailed usage of the GenRA. For instance, can ‘batch search’ give some bioactivity profile of chemicals?
Will you be able to separate out NAMs data from the animals’ toxicity data to look for similarities in specific MoAs of the chemicals? Dr Patlewicz responded that this is likely to be difficult to implement in the current version but exporting out specific fingerprint sets could be implemented in a future release. Participants are encouraged to submit feedback and enhancement requests on the GenRA home page by using the ‘Contact GenRA’ email link. Currently a user can select a particular fingerprint, and a search is performed to return the most similar analogues. By default, the number of analogues returned is 10 based on Morgan chemical fingerprints and prefiltered based on availability of ToxRefDB v2 data, however the number of analogues returned can be adjusted and the toxicity filter can be switched off. The underlying in vivo toxicity data supporting any prediction using GenRA is accessible using the Download file, however additional toxicity data is available elsewhere within the Dashboard. The Dashboard also provides the functionality to export out the bioactivity profile for a set of chemicals. Using the batch search and selecting the option (Associated ToxCast Assays) will download a file which depicts a heatmap of assay hitcalls for the selected chemicals. This download is referred to as an enhanced data sheet. Using this enhanced view, you can explore the data to identify trends in terms of where the activities are similar and dissimilar for sets of chemicals.
There was interest if the issue of metabolism in the bioactivity essays will be implemented in the dashboard. Dr Williams confirmed this work is ongoing, and Dr Patlewicz commented that in the context of similarity, metabolic similarities are often discussed. They have a proof-of-concept study ongoing in which they test the number of chemicals, and then look at using mass spectrometry to identify the metabolites that have been formed and how that corresponds to predicted metabolism from different software, with special attention to concordance with literature. Preliminary work in this study has been published [6]Boyce, M., et al., Comparing the performance and coverage of selected in silico (liver) metabolism tools relative to reported studies in the literature to inform analogue selection in read-across: A case study. Computational Toxicology, 2022. p. 100208. though another manuscript is in preparation that is more focused on the mass spectrometry analysis.
There was also interest in whether the Dashboard will anticipate a web service’s access to some functionalities for integration with ECHA computational capacity. Dr Williams mentioned an intensive rebuild ongoing for API at the moment that will allow it to connect to the data sources efficiently using the Dashboard.
Footnotes
- ^ Shah, I., et al., Systematically evaluating read-across prediction and performance using a local validity approach characterized by chemical structure and bioactivity information. Regul Toxicol Pharmacol, 2016 : p. 12–24.
- ^ Tate, T., et al., Repeat-dose toxicity prediction with Generalized Read-Across (GenRA) using targeted transcriptomic data: A proof-of-concept case study. Computational Toxicology, 2021: p. 100171.
- ^ Boyce, M., et al., Comparing the performance and coverage of selected in silico (liver) metabolism tools relative to reported studies in the literature to inform analogue selection in read-across: A case study. Computational Toxicology, 2022. : p. 100208
- ^ Helman, G., et al., Generalized Read-Across (GenRA): A workflow implemented into the EPA CompTox Chemicals Dashboard. ALTEX, 2019 : p. 462–465
- ^ Helman, G., et al., Generalized Read-Across (GenRA): A workflow implemented into the EPA CompTox Chemicals Dashboard. ALTEX, 2019. p. 462–465.
- ^ Boyce, M., et al., Comparing the performance and coverage of selected in silico (liver) metabolism tools relative to reported studies in the literature to inform analogue selection in read-across: A case study. Computational Toxicology, 2022. p. 100208.
5.5. Challenges of NAMs in the real world 1
Presentation of the Nordic Survey results
By Dr Hubert Dirven
Upon registration for the workshop, the participants were asked to complete an online survey. Dr Dirven presented the result from the survey completed by 22 participants.
The respondents, representing the Nordic competent authorities, reported to be working in Norway (36%), Sweden (32%), Denmark (23%) and Finland (9%). The majority (64%) had some experience in using NAMs in a regulatory context.
Most of the respondents reported some (86%) or high (5%) level of familiarity with using grouping and read-across, while 9% reported no familiarity with this. Furthermore, the respondents reported some (73%) or high (4%) level of familiarity with QSARs, while 23% had no prior experience. Among those reporting to be familiar with grouping and-read across, 91% had used QSARs in grouping and read-across, and 9% answered that they had used other NAM data for this purpose.
The survey identified that several QSAR tools have been used. The OECD QSAR toolbox and Epi win were used by 36% and 15% of the respondents, respectively, while others reported to have used Vega (6%), Opera (3%) and others (18%).
More than half of the responders reported to have some familiarity with the Tox21/ToxCast data for bioactivity data, and 14% had some knowledge of the CompTox Dashboard. The AOP and/or AOP wiki were familiar to 68% of the responders. The majority of the respondents (83%) were not familiar with the omics technology.
When asked about the main challenges in using grouping and read-across for regulatory decision making, 24% said that more training and hands on examples are required, 21% mentioned the lack of safety data on the groups of chemicals, and 17% reported that the data being scattered across many sources, including journal and publications, is challenging (Figure 3).
Figure 3. What are the main challenges in using grouping/read-across for regulatory decision making?
Finally, the participants were asked how they perceive the opportunities and/or challenges to include NAMs in their grouping/read-across. Several opportunities were identified, as illustrated in Figure 4.

Figure 4. What do you perceive are the opportunities/challenges to including NAM in your grouping/read-across?
Based on the survey results, Dr Dirven concluded:
- Good familiarity with grouping/read-across in regulatory decision making
- QSAR is the NAM that is most often used in grouping/read across
- A variety of QSAR toolboxes are used in the assessments
- Opportunities in using NAMS in assessments are identified
- Missing data or difficult to find data is often reported as a problem in grouping/read-across
5.6. Challenges of NAMs in the real world 2
OECD Survey on omics: results and considerations for follow-up activities
By Dr Magda Sachana
Dr Sachana presented the results from a survey on the use of omics in a regulatory context. In this study, where 45 replies were given, 44% were research scientists, 34% constituted risk assessors, policy scientist/specialist and 11% represented science and technology, advisors, chemists, academics. OECD members from several different countries were represented. Among them, 65% had some experience using omics data in a regulatory context and 29% had no previous experience, and the rest had a great experience.
To get a better understanding of the background of participants, they were polled on whether they were currently exploring the use of any omics data. 30-50% were using it for weighted evidence (49%), MoA (42%), priority settings (33%), grouping and read-across (42%), 18% of responders did not use omics data at all.
The OECD was interested to map and understand the challenges of using omics data in chemical risk assessment faced by the participants of the survey. “A lack of confidence in the data (here defined as lack of reliability, inability to link the omics data to endpoint or population level effect)” was most frequently identified as the main obstacle (ranked first by 29.5%). The most common response for the second most challenging obstacle (selected by 22% of responders) was “a lack of explanation as to when omics data is submitted for evaluation (i.e. why was the omics data generated, how reliable and reproducible is it, what does it mean/what difference does it make to the hazard and/or risk assessment)”. 26.3% of responders identified the third greatest challenge as “a lack of technical guidance on acceptable practices for omics data”. “The lack of confidence in omics data” was the most frequently selected response for the fourth most challenging obstacle. Among these “a lack of relevant examples (or case studies) to refer to” was mentioned. Finally, the most common answer for the fifth most challenging obstacle was “the lack of ‘in-house experts’ on omics to support regulatory programmes” (20% of responders).
The OECD also asked participants how one could increase confidence on technical aspects of the use of omics data in chemical risk assessment. “A linkage of omics data to endpoints used in regulatory decision making” was the most important criteria for 40.9% responders. “Having technical performance criteria (as part of acceptable practice)” and “the linkage of omics data to endpoints used in regulatory decision making” were tied as the second most important factor (with 24.4% of the respondents each). “A comprehensive review of case studies with omics data demonstrating sensitivity, specificity and reproducibility of data” was ranked as the third most important factor by 25.6% of the responders. “Having technical performance criteria in place (as part of acceptable practice)” and “standardised reporting templates for omics data” were tied as the fourth most important factor (25% of responders each). Finally, “having additional case studies demonstrating context of omics data use for difference decision-making scenarios” and “standardised reporting templates for omics data” were tied for the fifth most important with points (35.7% of the response each).
Certain actions could probably increase confidence in the use of omics data in chemical risk assessment (beyond the reporting frameworks). Ranked as most important (by 25.6% of responders) was “The development of an OECD guidance document(s) on application of omics data in regulatory decision making”. “The development of an OECD guidance document(s) on application of omics data in regulatory decision making” was ranked second most important by 20.5%. “The development of additional case studies demonstrating context of omics use for difference decision-making scenarios” and “incorporation of an omics section in the upcoming update of the grouping guidance” were tied as the third most important factor (each ranked that way by 18.4% of responders). “The identification of regulatory scenarios for which data from non-standard methods (i.e. omics data) can be used” was ranked as the fourth most important factor by 33.3%. Finally, 22.6% of responders felt that “a validation body or scientific advisory statement that supports the use of omics for chemical safety decisions” was the fifth most important factor.
The majority of the respondents (80%) agreed that MoA classification might be beneficial in their regulatory jurisdiction over the next 5 years. 71% felt that category formation to support biological read-across (RAx) might be beneficial in that time frame while only 43% stated that deriving a POD was likely to be beneficial.
There was also interest to know how the OECD could facilitate the future use of omics data in chemical safety assessment. “Provide training” was ranked as the most important by a majority (40%) of respondents, followed by “develop application reporting modules to report omics data, guidelines, technical, acceptable practices” which was ranked as most important by 32% of respondents. 12% of respondents ranked “coordinate and provide case studies” as the most important, 8% felt it was “describe validation/technical performance details. Only 4% felt that “Develop acceptable practices” was the most important and 0% responded that “provide guidance for use/interpretation” was the most important. When asked for the appropriate timeline for OECD work on additional omics projects, majority respondents indicated that this should happen in the short term and immediately (45%, 40%, respectively), indicating a strong need for this type of work.
Conclusions, remarks, and next steps
With this survey the OECD was able to map the biggest challenges for the use of omics in regulatory perspective and these were: 1. lack of confidence, lack of technical guidance, and lack of training for omics data. Indeed, the biggest tasks are to increase technical confidence. 2. develop linkage to regulatory endpoints 3. need to develop standardised reporting templates 4. lack of guidance documents and finally 5. lack of example case studies.
The next steps are to develop application modules (grouping and read-across, biomarker reporting frameworks), using a broad spectrum of case studies (i.e. look at omics in multiple cell types - complementarity for coverage of biological space; considering how to use POD and not tied to an apical endpoint in a regulatory context).
Under the application of omics two approaches could be followed, the pathway approach and the pathway agnostic approach. The pathway approach may integrate well with AOP frameworks and provide insight on the mechanism of action. A targeted pathway approach may limit sensitivity (i.e. may not be the most sensitive endpoint) while a pathway agnostic approach will provide a protective POD but may not provide a mechanistic understanding, and may not be needed for hazard identification.
For these activities the OECD webpage on omics was created (oecd.org/chemicalsafety/testing/omics.htm) and a recent paper was published [1]Harrill, J.A., et al., Progress towards an OECD reporting framework for transcriptomics and metabolomics in regulatory toxicology. Regul Toxicol Pharmacol, 2021. 125: p. 105020. .
Questions, comments, and discussion
It was pointed out that design omics study will be of great importance for omics (e.g. cell lines used, exposure system) and the question was raised as to whether the document already contains detailed recommendations? There is no recommendation yet, since this is still work in progress, however, what is known is that by using few cell types it is possible to predict the MoAs for unknown chemicals. Yet for known chemical MoAs special recommendations are required. For instance, if chemical A acts on a given receptor, in the design it has to be considered that the receptor needs to be present in the cell used.
It was stressed that omics is a great technology to measure response to stressors, however we should always think about the relevancy of the test systems (technology will never replace the relevance). Also, omics could be combined with in vivo systems to gain predictive power (i.e. successful applications of omics and 5-days rodent assays in US, omics and 28-day repeated dose toxicity assays in Japan). These results are quite impressive, because then you have an almost complete system.
There was a concern underlined the transparency of the omics data, the complexity of the data, difficulties to understand, time spent to conclude, and problems of transferring the knowledge to smaller institutions. The term ‘black box’ was used to recapitulate the concern.
As a response to this comment, OECD plans to address the issue of transparency, so that reporting will include elements that should allow all steps to replicate the results. It was also stressed that we do not need to understand all of the mathematics behind all the models. There are many standardised procedures that can be followed, and the field is matured, with >100000 papers on metabolics published in the medical field. We need to have clear reporting guidelines. It was also pointed out that once the regulatory agencies and the implementation of NAMs in regulation becomes more common or more adopted, we can rely on the private sector as well to gain the momentum in terms of utility.
Footnotes
- ^ Harrill, J.A., et al., Progress towards an OECD reporting framework for transcriptomics and metabolomics in regulatory toxicology. Regul Toxicol Pharmacol, 2021. 125: p. 105020.
5.7. Plenary discussion training needs
Prioritising the challenges identified - group discussion
By Dr Hubert Dirven
Small groups were formed to discuss what are the main challenges identified in using NAMs in regulatory assessment, and what should be prioritised. Discussions touched upon the need for knowledge (training), communication, confidence and guidance provided for future users of NAMs in regulatory contexts.
Training
Training needs were independently identified and highly ranked by all groups in this workshop. It was pointed out that similar workshops would facilitate training and dialogue between scientists and regulators. The importance of training the regulators was especially stressed, emphasising the major gap of knowledge for this group. More precisely the comments about training referred to: 1) how to understand the NAMs and interpret their results, 2) how to understand the importance of NAMs in the content of the regulatory and legal settings, 3) the need for case studies, for the validation and standardisation of NAMs (i.e. OECD reporting templates; NAMs useful for grouping and read-across, human relevance evaluation, MoA; challenges with negative read-across), 4) how to obtain a better understanding of how the data was generated, and 5) training/experience from primarily omics assays i.e. lacking the knowledge to interpret them properly, and therefore, to gain confidence in NAMs.
One group stressed the importance of setting future training goals and objectives. They also expressed some confusion over the end goal of using NAMs, in particular whether the intention was to support weight-of-evidence assessments, or to replace in vivo tests.
Communication
Need for more communication was indicated. Better dialogue between experts and regulators is necessary. It was suggested that ECHA could facilitate this communication. Practical training sessions with concrete examples of using NAMs would be beneficial. One obstacle was how QSAR data would be weighted. How solid does the evidence need to be for it to count?
Study design, confidence and relevance
Groups also pointed out that a proper design of the study is a necessary condition to ensure robustness of the data. Training is necessary to help regulators obtain a better understanding of how the data is generated and how the data were generated for specific applications. There should be very good guidelines describing specific processes to follow once the data is generated.
Surpassing black box and transparency
Other people also suggested that information from NAMs could represent a black box, and there will be a need to better understand how to navigate through the resulting data in order to know how to use it. Case study workshops would be very good for that purpose again. Others referred to the need for transparency in how the data has been derived. This also raised the need for more training in omics assays. The importance of transparency was also associated with the need for very stringent descriptions of the applied methodology. Some hesitation about confidence and prematureness were also expressed. There was also confusion from some members of the workshop about what we should be trained for, and they also wondered if these concepts were mature enough.
Another point was transparency. It is important that the regulators understand NAMs methods and how they can be implemented in the regulatory process.
New skills, in house experts necessary
There is a need for new complementary sets of skills in omics and toxicology for regulatory risk assessment. Detailed guidelines are also required for determining the relevance of NAMs (this can be very challenging, especially for omics derived NAMs).
5.8. Experience of ECHA in applying NAMs in a regulatory context
By Dr Tomasz Sobański
In his talk Dr Sobański presented the motivation of ECHA to enter the world of NAMs, and their experience of applying NAMs in a regulatory context. He presented the future vision of ECHA for speeding up regulatory work using these approaches. Both hurdles and gains from regulatory perspectives were thoroughly discussed.
New Approach Methodologies (NAMs) from ECHA perspectives
Dr Sobański extended the definition of NAMs presenting the ECHA perspectives. It includes in silico, in chemico, in vitro and in vivo methods that could enhance the pace of regulatory work. These will lead to more relevant decisions and will substantially contribute to the 3R principle (Replace, Reduce, and Refine). The main focus is still human health and environmental ‘endpoints’. He stressed the added value from NAMs comes from their high throughput and robustness, and from the increased understanding of toxicological MoAs they provide, which can be used to deliver appropriate protection levels for human health and the environment.
Ongoing activities in NAMs
ECHA is currently involved in multiple NAMs projects with the ultimate goal of translating these NAMs into regulatory applications. Among these, Dr Sobański mentioned ECHA’s participation in the APCRA initiative and the consortium case studies, its contributions to multiple EU research programmes (e.g. EU-ToxRisk, ASPIS cluster, PARC), involvement in specific tasks for the implementation of the EU CSS (using ToxCast/Tox21 assays and QSAR predictions under GMT) and its contribution to the development and adoption of Defined Approaches (e.g. skin sensitisation). ECHA is active in giving input to harmonisation and reporting. As an example, Dr Sobański mentioned its contributions to OECD Expert groups working on Transcriptomics and Metabolomics Reporting Framework (TRF & MRF) and QSAR assessment framework (QAF) to revise principles and establish an assessment framework. ECHA also takes an active role in the promotion of NAMs via training about NAMs for ECHA staff and committees, industry and scientific community, participation at various scientific and regulatory workshops and conferences and developing and maintaining tools and guidance. Finally, ECHA is active in the field of computational toxicology methods development (e.g. QSAR Toolbox).
Challenges in the hazard characterisation and assessment
Dr Sobański pointed out challenges in the hazard characterisation and assessment in regulatory contexts. In particular, he mentioned that although there are provisions in REACH for the promotion of the development of alternative methods for assessing the hazards of substances (Article 13: Animal testing as a last resort) there are also significant hurdles to be overcome. In particular, the information requirements in REACH (as well as classification criteria) refer to animal tests, and often to specific OECD in vivo test guidelines, indicated in the REACH Annexes.
Mapping experience of ECHA in development of NAMs in different project (regulatory perspective)
In the category of “current science using NAMs”, QSAR and read-across tools are used with the ultimate aim of replicating the traditional types of results. They give the same information to perform risk assessment, classification, and the labelling of chemicals. Dr Sobański stressed that if they are done correctly, they should provide us with equivalent information to be used in chemical regulation. Under the same category we can find a mechanistic understanding (i.e. AOP) which could deliver information to reduce uncertainty in risk assessment, classification, and labelling. However, there is still a serious knowledge gap about the relationship between molecular key events and the adverse outcome. Therefore we cannot address our concerns in a systematic way for the majority of the systemic toxicity “endpoints”.
Discussing the category, “new technology” Dr Sobański focused on omics. In principle, these approaches can cover the full spectrum of biological responses and promise to address human relevance (pathway relevance), however, they still need animals for confirmatory studies. In addition, the data produced are currently complex making interpretation difficult for regulatory bodies. This is a serious limitation of the new technologies in regulatory applications.
For ‘simple’ endpoints with local effects, the effort has been focused on in vitro and QSAR, with examples of successful outcomes (e.g. skin sensitisation), however, this is still impossible for complex, systemic endpoints. Existing NAMs and QSARs often fail to be used under the current regulatory framework as they are unable to provide the information required by the guideline studies (information of equivalence).
Criteria for regulatory acceptance
Dr Sobański further discussed what are considered the criteria of legal adaptations and the provision on risk characterisation in Annex I, section 6. Currently, in ECHA there is a consensus that an acceptable alternative method (NAM) should: a) identify the given hazardous property with comparable sensitivity and specificity to the respective animal test, b) enable classification of the chemical, and c) enable setting DNELs and PNECs, which implies that a test method or adaptation must provide strictly quantitative results (not only qualitative yes/no answers).
Criteria and the main challenges for regulatory acceptance of NAMs
Dr Sobański dived into discussing the main hurdles for regulatory acceptance of NAMs. The main challenge lies in the nature of the endpoints traditionally measured for regulatory processes. The spectrum of observed effects in higher tier endpoints (near adverse outcome pathway) is very broad (organ or body weight changes, biochemistry, histopathology, number of offspring, animals’ behaviour, etc.). In order to adapt to the information requirements, REACH is assuming equivalence in the levels of information. However, NAMs are not able to replicate all (or even provide equivalence) for this wide spectrum of effects and, even if theoretically possible, it won't be feasible due to costs. In addition, NOAELs (used to derive DNELs) and LOAELs which are used for hazard and risk assessment under REACH are relates to observed adversities. NAMs can provide a wide spectrum of biomarkers which might be predictive of adversity, but this is not accommodated under the current practice nor in classification and labelling criteria. Dr Sobański pointed out that even the most mechanistically driven endpoints, such as endocrine disruption or genotoxicity, are still based on adversity criteria.
Finally, an important hurdle, the majority of the current NAMs have not undergone formal validation and are not approved in the OECD test guideline programme, therefore, at the moment, they do not satisfy the principle of Mutual Acceptance of Data (MAD) agreed between the OECD countries.
Paradigm shift in regulatory risk assessment
Due to this hurdle, Dr Sobański concluded that because we have realised that alternative methods probably cannot directly replace animal tests (e.g. sub-chronic and reproductive toxicity) due to the inherited design of these tests, we should consider a longer-term “paradigm shift”.
This paradigm shift would imply that: 1) We anticipate that in vitro, in chemico and omics-based methods will develop and will be promoted in the future 2) We recognise that uncertainties are not only introduced by NAMs but they are also associated with safe values which are based on animal data 3) We accept that early indications of toxicity (e.g. biomarkers, and omics-data) can be used in regulatory decision making, provided that the legal provisions allow it 4) We find ways to use systemic concentrations instead of external doses, as the primary exposure metric 5) We initiate a comprehensive revision of the CLP criteria.
Building confidence for NAMs
While waiting for the paradigm shift, we could focus on grouping and read-across methods which are the most commonly used alternative methods [1]ECHA, The use of alternatives to testing on animals for the REACH Regulation. Fourth report (2020) under Article 117(3) of the REACH Regulation. 2020.. For instance, applying read-across correctly reduces the need for experimental tests because information on a similar substance (source) is used to predict the properties of another (target). A properly justified read-across can be used to fulfil REACH information requirements when Test Guideline studies have been used to generate the data for the source substance, and it is compatible with CLP. Unfortunately, while ECHA has advocated using grouping/read-across, it has had to reject the majority of read-across arguments due to lack of scientific rigour in defining groups of substances. NAMs could help in strengthening the read-across hypothesis and could be used as bridging evidence, however more discussion and guidance is needed to define how NAM data should be used.
Questions, comments, and discussion
It was stressed that all aspects discussed already were from the perspective of the specific regulatory application (so called “definitive hazard assessment”) where NAM evidence could be used directly to conclude on hazard and risk management. However, there are also other applications where NAMs can be used already like screening, prioritisation, or weight-of-evidence.
We should increase the scientific confidence in NAMs, one way of doing it is to apply molecular mechanistic data to support grouping/read-across cases. However, that would require an open discussion how NAM evidence can most effectively be used.
There is a serious gap in knowledge for AOP that should be filled with knowledge and data. At the moment we cannot address our concerns in a systematic way using AOPs.
Due to animal welfare considerations, we rarely have multiple data sets for the same hazardous property (endpoint) on a given chemical. This makes it difficult to quantify the reliability of in vivo data.
NAMs could be used as predictive tools to indicate the adversities with a certain probability and this needs to be addressed in the regulatory system, if we want to be successful in applying these methods.
Footnotes
- ^ ECHA, The use of alternatives to testing on animals for the REACH Regulation. Fourth report (2020) under Article 117(3) of the REACH Regulation. 2020.
5.9. Omics-based grouping/read-across (G/RAx): A case study with azo dyes
By Prof Mark Viant
In his talk, Prof Viant presented a read-across case study using a group of chemicals called azo dyes. The principal aim of study was to investigate the application of omics for grouping, irrespective of the biological test system and test substances, and to focus on developing a biologically-based grouping case study as part of a collaboration between ECHA and multi-omics experts.
Workflow of using omics for grouping
The main objectives of the presented grouping/read-across case study were 1) to select a target substance and series of potential source substances for grouping/read-across. This was done using an analogue approach; 2) to apply conventional approaches to form a grouping hypothesis; 3) to apply NAMs (in this case omics) approaches to substantiate the grouping hypothesis based on molecular mechanistic data using transcriptomics and metabolomics; 4) to map ToxPrint chemotypes onto the omics-based grouping to disentangle structural features of the molecules responsible for driving the biologically-based grouping; 5) finally, to conduct read-across to fill the data gap.
Classical grouping
Prof Viant used a target substance, an azo dye (Disperse Yellow 3; DY3), in comparison with six other source azo dyes. They used ECHA’s Read-Across Assessment Framework, and in this exercise the Daphnia model was used, with the endpoint (to read-across) chronic reproductive toxicity.[1]OECD, Daphnia magna Reproduction Test (OECD TG 211). 2018. This project used the QSAR toolbox to predict the group membership of the seven dyes using a range of different QSAR profiling approaches. Based on OASIS, ECOSAR 2.0, EPA and OECD chemical categories in the QSAR Toolbox, they concluded that DY3 falls outside of the other two categories formed from the rest of the dyes used in this exercise. They also concluded the target dye was closer to the subgroup of Sudan dyes.
Next, they grouped chemicals using a structure dissimilarities matrix of the molecules (based on ToxPrint chemotypes and Tanimoto distance matrix) and concluded this analysis with hierarchical clustering. DY3 was found to be quantitatively more similar to Sudan 1 (S1) and Sudan Red G (SRG) than the rest of the dyes used, consistent with the QSAR profiling.
Mechanistic grouping using multi-omics
The gene expression levels and metabolomic profiles were measured for 3 different doses and 3 time points. Prof Viant integrated these two types of omics data for multi-omics grouping of the chemicals. This was based on hierarchical clustering, supplemented with an uncertainty analysis of the formed clusters to ensure that the grouping was reliable. In this multi-omics analysis, the target DY3 was most similar to S1, which is the reason it was chosen to be the source substance for later read-across. The individual omics approaches produced similar grouping results, however they had lower confidence. This suggested a multi-omics approach for grouping would enhance confidence.
To get a better mechanistic understanding of the omics results, they presented molecular pathway enrichment analysis for the target and the 2 Sudan dyes that showed the closest biological responses. The biological response was presented as a heatmap and ranked by top responsive pathways. Here, presented ‘cellular stress’ and ‘injury’ pathways could be identified. By mapping the ToxPrint chemotypes of all structural features onto the multi-omics-based grouping, they predicted the hydroxyl group in the dyes caused the toxicity, i.e., that the biological response-based grouping is being driven by this particular structural feature. In this way the biology and chemistry map together consistently to make a more rigorous understanding of what is happening in this system.
Read-across validation
An analogue approach with single target substance and single source substance was used for read-across (read-across from a single source substance to a single structurally similar target substance). Read-across to predict toxicity was done from S1 to the target DY3. For this purpose, both dyes NOEC and LOEC were measured. NOEC was identical, and LOEC were very similar for both molecules in the Daphnia model, experimentally validating the multi-omics-based grouping.
Conclusions
There is a need to establish workflows that enable molecular (omics) mechanistic data to be used alongside conventional structure-based grouping approaches (as presented in this case study). The uncertainties in conventional approaches for grouping could be reduced significantly by incorporating omics data, in particular whenever the conventional structure-based approaches show inconsistencies. Even for the case where structure-based methods provided consistent grouping, there can still be doubt as to whether these methods are proposing the correct grouping hypothesis. Therefore, further confidence in the grouping hypothesis should be sought by incorporating biologically-based grouping alongside structure-based methods routinely. A further benefit illustrated by the case study is that by mapping ToxPrint chemotypes onto the omics-based grouping, predictions can be made about which part(s) of the test substances are driving the biologically-based grouping.
Footnotes
- ^ OECD, Daphnia magna Reproduction Test (OECD TG 211). 2018.
5.10. Experience ECHA in applying NAMs in a regulatory context
Use of omics approaches - text analytics case from ECHA
Generally, there was little occurrence of the words ‘metabolomics’ in IUCLID dossiers. Most of the entries were related to physicochemical properties toxicokinetics, including metabolism. In total, there were only a few tens of unique registered substances.
In addition, it is important to consider the data analysis methods and interpretation. Examples include principal components analysis, clustering methods and correlation-based methods. It was noted that the mechanisms elucidating the MoAs were not always described. The information presented in the dossiers does not, therefore, always use the full potential of omics data, because it does not consider the mechanistic aspect of the information.
Potential use of omics and regulatory hurdles
Dr Bouhifd pointed out that to use such data as a read-across adaptation in REACH, the requirements set in annex XI, 1.5 should apply. These requirements have been listed and it was stated that even if a metabolomics study provides information to support a hypothesis for read-across and that the read-across justification might be plausible, issues could be raised in relation to the adequacy of the supporting information and also to the availability of supporting information (e.g. bridging the data with the substance as in screening studies).
NAMs and regulatory requirements
Conclusions
- There is still very limited submission of omics data in general. A question for discussion is whether this is due to the lack of guidance or the reluctance by risk assessors and managers to use this type of information.
- Steps in the data analysis that lack transparency must be removed.
- The reporting is an important aspect to be addressed.
- Detailed descriptions of the pathways that link adversities need to be provided, otherwise the promise of mechanistic omics in toxicology risk assessment is not really fulfilled.
5.11. Variability in repeat dose toxicity studies
By Dr Katie Paul Friedman
Implications for scientific confidence in NAMs
In her talk Dr Paul Friedman focused on the acceptability of NAMs for risk assessments. Expectations are that NAMs must provide information of equivalent or better quality than traditional animal models. That brings the question, how well do the animal tests actually perform? We need to know what the variability of traditional effect levels might be in order to know one aspect of assessing acceptable values from a NAM.
Dr Paul Friedman and her research group used the database ToxRefDB v.2.0. to evaluate variability in traditional data for more than 1000 chemicals and more than 5000 studies. Based on the study descriptors included in the database, they developed statistical models of the variance in effect level values. The results from 28 different statistical models, suggested that the variance that can be explained by study descriptors lies in the range of 55-73% [1]Ly Pham, L., et al., Variability in in vivo studies: Defining the upper limit of performance for predictions of systemic effect levels. Comput Toxicol, 2020. 15(August 2020): p. 1–100126.. The estimates corresponded well with previous work in this area. In other models of large datasets of in vivo repeated dose toxicity data, typically 50-70% of the variance have been explained [2]Helma, C., et al., Modeling Chronic Toxicity: A Comparison of Experimental Variability With (Q)SAR/Read-Across Predictions. Frontiers in Pharmacology, 2018. 9. [3]Mazzatorta, P., et al., Modeling oral rat chronic toxicity. J Chem Inf Model, 2008. 48(10): p. 1949–54. [4]Toropov, A.A., et al., CORAL: model for no observed adverse effect level (NOAEL). Mol Divers, 2015. 19(3): p. 563–575. [5]Toropova, A.P., et al., The application of new HARD-descriptor available from the CORAL software to building up NOAEL models. Food Chem Toxicol, 2018. 112: p. 544–550. [6]Toropova, A.P., et al., QSAR as a random event: a case of NOAEL. Environ Sci Pollut Res Int, 2015. 22(11): p. 8264–71. [7]Veselinović, J.B., et al., The Monte Carlo technique as a tool to predict LOAEL. Eur J Med Chem, 2016. 116: p. 71–75.. Dr Paul Friedman noted that acceptable NAM accuracy should be informed by more realistic expectations. A primary conclusion from this work was that the variability in in vivo toxicity studies limits predictive accuracy of NAMs that use these data as reference or training information. Given the understanding that uncertainty in in vivo data used in training will lead to uncertainty in NAMs, Dr Prachi Pradeep and colleagues developed a QSAR for repeated dose toxicity using estimates of variability in animal data to construct a POD distribution [8]Pradeep, P., K. Paul Friedman, and R. Judson, Structure-based QSAR models to predict repeat dose toxicity points of departure. Computational Toxicology, 2020. 16: p. 100139..
Ongoing work in Dr Paul Friedman’s group is now focused on understanding the variability in organ-level findings in repeat dose studies. This analysis focuses on treatment related changes in specific endpoint-related targets (e.g., any effect on the liver). Preliminary result suggests that within species there was 65-90% concordance in finding any effect in the endpoint related target tissue. Dr Paul Friedman thus suggested that the upper limit on accuracy for a NAM to predict the same target organ might be 60-90% depending on the model. This research also includes an analysis of the concordance of organ-level findings between repeat dose studies of different duration. If a substance failed to produce effects in a target organ at 90-day (subchronic study), what are the odds there would be a positive finding in a 1–2-year study (chronic study)? Approaching this question, a positive finding was defined as any gross or histopathologic change or associated hormones or clinical chemistry in the target organ. Their work demonstrated that a positive finding in the subchronic studies tended to indicate a greater likelihood of a positive finding in the chronic studies. These findings suggested that a POD in a target organ derived from a subchronic study (particularly for liver and kidney with the largest datasets) is most likely protective for any chronic finding on the basis of dose (ignoring the level of adversity of the findings).
Finally, Dr Paul Friedman previewed work in progress on an adjustment factor that might be needed when doing NAM based assessment of repeated dose toxicity, using in vitro assay data and a high-throughput toxicokinetic modelling approach to estimating in vivo doses that would be bioactive in liver or kidney. Currently, there was only enough data for kidney and liver in the ToxRefDB V2.0 to make the comparison, but the PODs based on bioactivity generally provided a protective POD at the organ level.
Footnotes
- ^ Ly Pham, L., et al., Variability in in vivo studies: Defining the upper limit of performance for predictions of systemic effect levels. Comput Toxicol, 2020. 15(August 2020): p. 1–100126.
- ^ Helma, C., et al., Modeling Chronic Toxicity: A Comparison of Experimental Variability With (Q)SAR/Read-Across Predictions. Frontiers in Pharmacology, 2018. 9.
- ^ Mazzatorta, P., et al., Modeling oral rat chronic toxicity. J Chem Inf Model, 2008. 48(10): p. 1949–54.
- ^ Toropov, A.A., et al., CORAL: model for no observed adverse effect level (NOAEL). Mol Divers, 2015. 19(3): p. 563–575.
- ^ Toropova, A.P., et al., The application of new HARD-descriptor available from the CORAL software to building up NOAEL models. Food Chem Toxicol, 2018. 112: p. 544–550.
- ^ Toropova, A.P., et al., QSAR as a random event: a case of NOAEL. Environ Sci Pollut Res Int, 2015. 22(11): p. 8264–71.
- ^ Veselinović, J.B., et al., The Monte Carlo technique as a tool to predict LOAEL. Eur J Med Chem, 2016. 116: p. 71–75.
- ^ Pradeep, P., K. Paul Friedman, and R. Judson, Structure-based QSAR models to predict repeat dose toxicity points of departure. Computational Toxicology, 2020. 16: p. 100139.
Conclusions
Construction of NAM-based effect level estimates that offer an equivalent level of public health protection as effect levels produced by methods using animals may provide a bridge to major reduction in the use of animals as well as identification of cases in which animals may provide scientific value. Existing QSAR for repeat dose POD may be informative for rapid workflows and is in progress to support best practices for estimating in vivo PODs at the organ level.
Questions, comments, and discussion
After her presentation, Dr Paul Friedman was encouraged to elaborated more on prediction intervals. She said that the prediction interval cannot be smaller than the estimated variance in the reference data, however it can be bigger. If a NAM adds variability to a response, you expect that the prediction interval increases. Because the minimum prediction interval is based on the variance in the training data and not on the variance contributed by some NAM, Dr Paul Friedman argued that one should be careful to call it the minimum prediction interval.
An obvious limitation, however, is the limited availability of replicate study data. Some repeats of single study types for a given chemical are available, but it is unlikely that this dataset will grow because the studies are seldom replicated in the regulatory framework, at least not in Europe where it is more common for registrants to work together in consortia to submit a study. Rerunning a study will most certainly affect the study-level POD, because there will be inherent variability in the performance of these studies, and that is very much appreciated by industry toxicologists.
Dr Paul Friedman was also asked to elaborate more on use of animal data versus human data in benchmarking NAM performance. She argued that comparison of NAMs to animal data is only one tool in the toolbox. When there are regulatory drivers behind the use of animal models, it is challenging to have any human data to evaluate NAM performance. As more NAMs are developed, efforts to develop NAMs that measure biomarkers that can and/or are measured in humans would facilitate more comparison of NAMs to human health indicators.
5.12. Plenary discussion
One of the participants was arguing, that when we are discussing NAMs we are pointing in many different directions. Particularly with respect to the time perspective. Some are expecting NAMs to be ready to replace in vivo testing tomorrow. ECHA, on the other hand, gives the impression that it will take an entire new legislation in the EU before NAMs can be used. How should we address and communicate this gap?
Several participants followed up on this.
The chemical strategy for sustainability does include the revision of the REACH regulation, and the policy makers are discussion new options for the amendment of the standard information requirement.
We already know that omics as a technology is useful, and maybe we do not have to wait for a paradigm shift to amend this information into test guidelines. It was suggested to introduce measurements and molecular data into the very same tests that are already being conducted.
Another point in the discussion was to avoid ending up in the same situation as with the cosmetic regulation, where animal studies were banned before the regulatory framework was ready.
And what about the economic aspect, moving away from expensive animal testing like the extended one generation reproductive toxicity study? From the regulatory side it has probably been a hard drive to use NAMs. But that is a difficult question and should be directed to the Commission.
Historically, when new technologies are adopted, the process is typically long and painful. The implementation of NAMs can be compared with the implementation of using DNA as evidence in criminal justice system, it did not happen overnight. We are now in the middle of the process of testing the technology to see how far we can take it.
The last contribution to the discussion was related to the inherent properties of chemical substances. We already know that we do not want to expose ourselves to bioaccumulating, persistent or genotoxic substances. Maybe there is not always a need to demonstrate or predict the adversity? If toxicologists came together, they would easily come up with a list of basic chemical properties that NAMs can measure very well.
5.13. Grouping and read-across case studies from EU-ToxRisk
By Dr Sylvia Escher
In her talk, Dr Escher presented case studies of employing grouping and read-across methods from the EU-ToxRisk project. The studies were designed to provide experience in using NAMs in multiple aspects of regulatory decision making. She touched upon how important the relevance of different NAMs models is, and how we can best integrate NAMs to come to conclusions in human risk assessment.
Paradigm shift for human Next Generation Risk Assessment (NGRA)
Dr Escher started her talk by presenting a common vision for next generation human risk assessment, where hazard characterisation identification (currently done using animal tests) could be replaced with batteries of in vitro tests and NAMs (such as multi-omics, in silico, organoids, grouping, AOPs, in vitro PBPK, qIVIVE, or human models). This would also require the development of Integrated Approaches to Testing and Assessment (IATA) and further extrapolation to human safety assessment. She pointed out the urgent need to demonstrate the relevance of different NAMs models, and how we can best integrate these conclusions into regulatory implementation.
NAMs opportunities and challenges
The opportunities of NAMs appear obvious and include areas such as the improvement of our mechanistic understanding of chemicals, human specific models, and the 3Rs principle (the replacement, reduction, and refinement of animal testing). However, challenges such as the need to implement and assess the robustness and reproducibility of assays, sensitivity and specificity, predictivity of these new assays, data analysis and integration (for new assessment strategies) still need to be addressed. There is also a need to further bridge the gaps in regulatory implementations such as a limited experience with NAMs and its multidisciplinary nature requiring different expertise. Special attention is now needed to assess the uncertainty of individual NAMs and combined approaches, NAM, fill the knowledge gaps, define relevance of NAMs and IATAs for toxicological endpoints and the scope of NAM testing, and find strategies for dealing with conflicting evidence.
Using NAMs case studies
The aim of the presented case studies was to demonstrate the integration process of NAMs for a human risk assessment for ‘repeated dose toxicity’ and ‘reproduction toxicity’, within a read-across assessment.
Read-across assumes a structure-activity relationship (SAR), so that often compounds with similar structural and physico-chemical properties are the starting point of the assessment. Within the read-across evaluation, convincing evidence has to be provided about shared toxicodynamic and kinetic properties within the grouped compounds to learn about a molecule of interest [1]Ball, N., et al., Toward Good Read-Across Practice (GRAP) guidance. ALTEX, 2016. 33(2): p. 149–66..
Target substances are often lacking information about ADME from their source components (especially for in vivo components) to extrapolate to the target chemical, and these are often estimated from physicochemical properties only. This is a drawback and NAMs data might be a goldmine to support the read-across assessment to both fill these data gaps and to gain predictive power. To improve that, the inclusion of compounds with interesting structure features will be useful for expanding the repertoire (i.e. investigating size chain properties simultaneously to the position of a group of interest in a carbon chain), and not being limited to the ana|lo|gues with only in vivo data points. In vitro ADME data allows for a greater understanding of kinetic properties that can be extrapolated to humans. NAM data can also provide mechanistic information and might thus be useful to support the assessment of shared toxicodynamic properties between source and target compounds.
Read-Across supported by NAMs - Systemic toxicity, lead effect liver steatosis
In a given example, one chemical (2-ethyl pentanoic acid) from a group of aliphatic, branched carboxylic acids (n=13) was used as a target in read-across using NAMs. Among the source components, only some of the chemicals had in vivo data available, and the compounds with high similarity scores gave conflicting effects (liver steatosis observed for 3 chemicals versus no in vivo effects at the highest tested dose for 2 chemicals). The EU-ToxRisk case study working group measured batteries of assays that were related to the different Molecular Initiation Events (MIEs) from a large AOP network on liver steatosis extracted from AOP wiki and other peer-reviewed publications. In addition, other unrelated MIEs and Key Events (KEs) were measured to cover a broader mechanistic space. Selected MIEs/KEs, (including lipid accumulation in cells after 24h, 72h and 10 days of exposure to the chemicals) were measured. They also supplied transcriptomic data from HepG2 cells exposed to different concentrations of the analogues tested. Dr Escher mentioned that the AOP wiki was supplemented with MIEs/KEs measured with EU-ToxRisk and KEs found in literature (testing scope was discussed).
Based on transcriptomic profiles, the biological similarities of the analogues were calculated using clustering methods, and the gene expression was visualised as a heatmap. As carboxylic acid doses increased, the expression patterns started to emerge, and two distinct clusters were formed (long-chain chemicals and short-chain chemicals) based on the similarity of the gene expression patterns. Also, the negative control (rotenone, a molecule with a distinct structure and different known MoA) did not cluster with any of these carboxylic acid compounds. The gene profile of carboxylic acids differed from rotenone. Also, the activation of early MIEs and KEs belonging to the AOP on liver steatosis as well as the late KE lipid accumulation in different human liver cells showed the same picture, able to divide chemicals for those that cause lipid steatosis. MIEs and KEs not related to the AOP did not show a consistent picture for the tested analogues.
In this exercise, researchers were able to classify/divide compounds into 2 groups and to conclude that analogues with longer side chains are more active and cluster together. The supplied results from MIE testing were put through a Dempster–Shafer selection procedure (decision theory based on Bayesian mathematics) to choose those MIEs that were most useful in classifying which chemicals cause steatosis [2]Escher, S.E., et al., Integrate mechanistic evidence from new approach methodologies (NAMs) into a read-across assessment to characterise trends in shared mode of action. Toxicol In Vitro, 2022. 79: p. 105269..
Footnotes
- ^ Ball, N., et al., Toward Good Read-Across Practice (GRAP) guidance. ALTEX, 2016. 33(2): p. 149–66.
- ^ Escher, S.E., et al., Integrate mechanistic evidence from new approach methodologies (NAMs) into a read-across assessment to characterise trends in shared mode of action. Toxicol In Vitro, 2022. 79: p. 105269.
NAMs can be used for ADME
The NAMs data were also used to investigate the bioavailability of different compounds (one of the ADME priorities). Only one of the chemicals had pharmacokinetic data (plasma concentration) in humans. The human in vivo ADME data were used to characterise the uncertainty of the PBK model. They used in vitro clearance data from primary human hepatocytes data and the predicted plasma protein binding values to parametrise PBK models with compound specific data.
The target chemical (2-EPA) activated the MIEs and KEs belonging to the investigated AOP. It also induced lipid accumulation in different liver cells. The results illustrated a shared MoA of chemicals. This shows that early MIEs/KEs can be used to provide information about similar modes of action (relevance of NAMs), while in vitro assays can be used to show trends in toxicokinetics (ADME). PBK simulations for all analogues identified a trend for increasing clearance and thus decreasing systemic exposure, with decreasing side chain length. As read-across aims to provide a POD for risk assessment, the PBK simulations can be used for in vitro to in vivo extrapolations to derive a human equivalent dose.
Computational modelling - Receptor Docking Modelling
NAMs are also useful in computational modelling approaches (i.e. receptor docking). Dr Escher gave an example of such an approach from the EU-ToxRisk project, related to the identification and characterisation of parkinsonian hazard liability of the chemical deguelin by an AOP-based testing and read-across approach. Degueline and rotenone (previously known from its toxicity) share similar pharmacophores. The identical docking posing of rotenone and deguelin into a crystal structure of Complex-I, causes the inhibition of this enzyme and manifests symptoms resembling parkinsones. They can inhibit molecular initiation events and they share the same toxicodynamic properties.
Conclusions
Using different case studies, including complicated AOPs networks, we do not need to test all the MIEs to come to the decisive conclusions. Dr Escher postulated that it is recommended to test a good representative number of those early key events and aim to show that they share the same MoA. Dr Escher also suggests using one of the later key events for the in vitro extrapolation to predict a POD in the risk assessment. Finally, decision making using Bayesian approaches can be a good way to interpret data.
NAMs come from multidisciplinary approaches, so we need many different types of expertise to overcome the challenges for the regulatory implementation. NAMs use and produce different systems and many types of data such as: in vitro, organoids, high-throughput, high content analysis and omics so, to be able to use it, we need various and different expertise. In addition to toxicology knowledge, omics experts, data infrastructure, data sharing, AOPs, and modelling Bayesian networks are needed.
Excellent science must come close together to the human risk assessment. Case studies based on a clear hypothesis have shown to be an excellent tool to bridge this gap. Standardised in vitro assays that are validated across labs with their performance are required to ensure that the test systems are robust and reproducible.
Questions, comments, and discussion
These studies opened discussion on how to integrate NAMs data into regulatory risk assessment practises in the future. Owing to the batteries of resources, background needed, and the multidisciplinary competence of the people involved, the participants wondered, what is actually the potential of using AOPs network NAMs in the near future? Is this feasible to happen for any endpoint? Dr Escher pointed out that we have different options already now, but more relevant AOPs need to be developed. She sees this as feasible.
Prof Viant commented that the way Dr Escher describes NAMs fits very clearly to broad pictures of the workshop, especially how NAMs should be used. This included the way AOPs were used in these studies, the way in which molecular key event biomarkers are used, the contrast between targeted and untargeted assays, and the way in which these can actually be called upon for different tasks. He pointed out that the gene expression data provided information about the AOPs upstream of the molecular initiating event.
Prof Viant wondered why Dr Escher made a recommendation to also test a key event close to the apical endpoints, as the benefits were not obvious, and may potentially be disadvantageous. He expressed a concern that doing so could reduce specificity. Dr Escher pointed out that for a large AOPs network, this can be advantageous, for some of the assays that are very sensitive and activate on very low doses compared to other assays. This is especially true in the case where we have many events and wish to predict an adverse outcome that is many events away. Using a key event that is close to the apical point can be useful as it will more likely also progress to the adverse outcome once activated. Therefore, it is advisable to include a key event that happens later in the AOP into the testing approach and to use its benchmark concentration to predict a POD for risk assessment. She stressed that this finding form the EU-ToxRisk case studies needs to be further explored in future to learn about its general applicability.
5.14. ECHA’s experiences with NAMs in the assessment of chemical groups
By Dr Tomasz Sobański
In this talk, Dr Sobański explains the rationale for setting up the Integrated Regulatory Strategy (IRS) in ECHA. He explained how the chemicals grouping supports the IRS to speed up regulatory actions and better manage their risks. He also gave examples of different workflows in ECHA.
The motivation of Integrated Regulatory Strategy
ECHA noticed that in the past, the various REACH and CLP processes were slightly disconnected at the operational level. For example, ECHA noticed that while performing a compliance check, not all consequences for CLP regulation were considered. To improve this, ECHA set up the Integrated Regulatory Strategy (a harmonised and holistic way of looking at all REACH and CLP processes). In addition to this integrated way of working through various processes ECHA also introduced the grouping concept to set up a more efficient way to deal with the substances within the groups rather than as individual cases.
For IRS, grouping was proposed on similar structures, features or properties and not necessarily based on their hazard. The grouping concept would work to find substances that require regulatory action. Such grouping could facilitate decision-making about the safety of the group, map progress, speed up follow-up, or the prioritisation of substances for testing. Finally, it would improve understanding of the need for regulatory actions. Work on groups is an informal preparatory task for the official regulatory processes and it does not aim to replace anything, but to improve the connections between legally binding decisions, consultations, and official processes. This strategy focuses on improving efficiency, reproducibility and the consistency of decision making, rather than assuming a uniform pattern of the hazard properties among the members.
NAMs are used in an iterative way for assessment of regulatory needs
For each group initiated by the GMT there is an assessment of regulatory needs performed by hazard and risk assessors. Experts produce a report with an overview section, the need for regulatory risk management (RRM), and a table showing the proposed immediate next actions and foreseen regulatory needs.
First, an assessment is done at the screening level via hazards profiles, using exposure profiles, group boundaries, potential for substitution, and the initial assessment of regulatory needs. This allows immediate action and expects further regulatory actions for the whole group, (sub)group, or for individual substances.
Then, data generation is proposed, if necessary although in some cases, risk management is already possible. The assessment of the read-across/category approach is only done during official processes (e.g. a compliance check). Finally, the depth of assessment and knowledge on substances will increase in further iterations. In this iterative assessment, NAMs mainly contribute to the assessment of hazard and, potentially, to the assessment of group boundaries.
The NAM input for ECHA processes
ECHA developed a multi-layer concept to interact with different regulatory processes. QSARs and NAMs can be considered as an input to ECHA processes at 3 different levels, with trade-offs in robustness and throughput compared to specificity of the applications.
Level 1 consists of a library of well characterised modules (NAM datasets and QSARs) which can be used to predict various properties of chemicals. Each module can be characterised by predicted property, type of output and its performance. This set of predictive tools and external datasets is used for building specific workflows for Level 2 and 3 (e.g. toolbox profilers, QSARs predictions, Tox21 and ToxCast datasets).
Level 2 considers streamlined applications (or workflows) for specific needs, which are of a repetitive nature. They are combining inputs from Level 1 modules to derive expected outcomes (e.g. predicted concern). Semi-automated scenarios to address specific needs can then take place, (e.g. input to a team of experts). These often require a combination of predictive tools, data and scientific knowledge. The workflows address the most common problems encountered during this. The workflows are not meant to provide definitive conclusions, they are designed to provide some additional information to assist hazard assessors and to allow them to make more confident and successful decisions (supporting evidence and supporting confidence).
Level 3 considers detailed and defined NAM and QSAR expertise to support ECHA needs. The problems addressed at Level 3 are all unique and require unique solutions. These could be ad hoc requests where QSAR and NAMs expertise or other expert consultations are needed (e.g. Compliance check under dossier evaluation, Substance Evaluation, and GMT). Assessment of technical equivalence for biocides, and input to projects like PARC may also be requested as level 3 input.
Workflows for Group Management Team (GMT) as an example of Level 2
Currently, 3 workflows are implemented (PBT/vPvB, ED screening, and metabolic profiling). These are now in the piloting phase. In addition, the following workflows are under development: B screening for ionisable substances, TK profiling, and in vivo alerts for CMRs.
Persistent, bioaccumulative and toxic chemicals (PBTs) screening workflow
A PBT screening workflow was implemented as a profiler tool in the QSAR Toolbox. The aim of this screening workflow is to screen for substances with the potential for P/vP, B/vB or T based on experimental data and QSARs predictions. The profiler was calibrated using the REACH guidance on PBT assessments. This workflow covers: a) applicability domain checks b) applicability domain evaluations inside the ECHA PBT screening profiler c) assessments of the relevant metabolites. Users are able to combine the ECHA PBT screening profiler with other OECD QSAR Toolbox tools.
Endocrine disruptors (ED) screening workflow
The ED screening workflow integrates predictions from multiple tools (e.g. VEGA, OPERA, US EPA CompTox Dashboard, Danish QSAR database platform – data from Endocrine Disruptor Screening Program in the 21st Century- EDSP21 and Derek in vivo alerts). It covers EAST modalities, in vivo alerts which might be related to ED, and supporting evidence from AhR, PXR, CAR, CYP3A4 predictions/data. At the moment, the final integration of evidence is done manually to gain confidence in the tools and refine evidence integration logic. Once the algorithm is fully developed there will be the possibility for higher-level automation.
ED screening for GMTs uses the hierarchy for evidence and evidence integration rules.
Here the rank is as follows: ToxCast ER/AR models (high confidence), multiple EDSP21 assays (high confidence), single EDSP21 assays (medium confidence), consensus QSAR models (medium confidence), single QSAR models and alerts (low confidence).
ADME and TK profiling often addressed in ECHA
These can address whether the metabolic profile of substance A is similar to substance B. It also can check the possibility that a substance of interest could be metabolised to a substance of concern (work on BPx groups). TK properties (such as bioavailability, T1/2, Cmax, AUC) of the substance(s) can also be investigated.
Final conclusions and remarks
- ECHA’s Integrated Regulatory Strategy aims to ensure that REACH and CLP processes are aligned, coherently implemented, and that they support authorities in identifying and addressing substances of concern as quickly as possible.
- The aim of Integrated Regulatory Strategy machinery is to 1. efficiently select (groups of) substances that raise potential concern. 2. generate the information needed to assess safety to address remaining concerns. 3. ensure appropriate and timely risk management by all actors (authorities and industry). 4. facilitate improved communication on safe use in the supply chain.
- The benefit of the grouping is that it treats related substances consistently, targets the right substances at the right time, pools information which may allow faster action (despite data gaps), and increases predictability of authorities’ actions. It constitutes preparatory work to support REACH and CLP processes.
- NAM input is used as supporting evidence for making conclusions on GMT hazard profiles, mainly to disregard MoAs concerns or to trigger the need for the generation of additional data.
- Workflows are designed to have high specificity at the cost of lower sensitivity.
5.15. EU-ToxRisk Case study 11
By Prof Bob van de Water
The case of a high-concern toxicity profile screening - EUToxRisk
Prof van de Water presented overarching goals and methods from the EU-ToxRisk project. This project uses varying complexities of data, and throughput strategies to establish the test systems toolbox. He stresses that omics approaches could deliver mechanistic insights and might be more sensitive at prioritising the chemicals than other approaches. He also gave many examples of other assays from which toxicological information could be extracted and used successfully for the classification. This work requires integration and is still ongoing.
Vision of EU-ToxRisk
EU-ToxRisk envisions delivering solid and pragmatic read-across procedures that will incorporate a mechanistic understanding, supported by toxicokinetic knowledge. It also aims to establish the starting point (ab initio) for hazard and risk assessment strategies for chemicals of scarce information (i.e. ‘low tonnage’ chemicals).
Prioritisation strategies could be supported by NAMs
The EU-ToxRisk aims at defining the compounds that fall in the category of high exposure and strong biological effects, or at lower exposure with high potency that could seriously perturb human biology. For this purpose, it is necessary to build a performance matrix, with a high confidence for prioritisation. Prof van de Water dives into the questions of the best way to use NAMs data to identify chemicals of concern for human health, how to define high exposure and strong biological responses, and whether we can use high throughput methods for NAMs.
One of the main hypotheses of EU-ToxRisk is that high throughput NAMs can properly rank ECHA substances based on mode-of-action in relation to toxicity to reproduction or endocrine-disrupting. It is also hoped to be able to rank the overall potency of the perturbations to human biology. He argued that, if that is the case, these could be added into the prioritisation toolbox of ECHA chemicals for further testing.
The main objectives are to 1) demonstrate the overall feasibility to identify substances likely to have a high toxicity profile based on high throughput NAMs hazard information. 2) use hazard information based on qualitative and quantitative mechanistic mode-of-action assessment in both in silico approaches and in vitro human test systems.
Tuning NAMs with ECHA in vivo data
In order to tune the NAMs, EU-ToxRisk complemented some of the ECHA in vivo data with NAMs approaches for the chemicals of interest. Their training set contained the available pools of chemicals with different potency (low, middle, and high toxic compounds with in vivo safety data), that were selected from a list of high tonnage ECHA chemicals. Using the toolbox of EU-ToxRisk, both in silico and in vitro NAMs, with a focus on high throughput, they tried to classify compounds based on their toxicity and potency status from the list. The data set consisted of the substances based on Annex 8 compounds (10-100 tonnes) that had 28-day reproduction and basic mutagenicity test data. This was to classify if substances were toxic or not in 90-day experiments.
Toolbox for EU-ToxRisk
This project uses various toolboxes, and, among these, the following were implemented: High Content Imaging (HCI) platform, CALUX assays, HepG2 BAC-GFP cellular stress reporter assays, LUHMES neurite outgrowth assay, and targeted TempO-Seq (Templated Oligo-Sequencing) sequencing. These tools were applied on different cell types.
HCI is a powerful microscope-based tool to run high-throughput toxicity assays. This can be done, for instance, using specific antibodies against a protein of interest, to track their presence and location in cellular compartments. Protein overexpression assays (i.e. CYP) could provide information on whether the toxicity MoA is related to the metabolism. The key message was that we need to take metabolic activity for the compounds of interest into account as well when assessing toxicity.
CALUX assays are based on the signals from luciferase activity that depend on a particular reporter that is incorporated into the cells (i.e. mostly nuclear hormone receptor activation and also several cellular stress response pathways). These assays can deliver benchmark concentration (BMC) values.
HepG2 BAC-GFP cellular stress response assays constitute another panel with a green fluorescence protein (GFP) fusion being used as a signal reporter to detect different points (i.e. oxidative stress, DNA damage, inflammatory stress or heat shock protein). Using live cell imaging these assays allow analysis of responses at the individual cell level. The assays have been licensed to Toxys and marketed as the ToxProfiler assay).
LUHMES cells are used in a neurite outgrowth assay and applied in imaging-based phenotypic assays where biological neurological perturbations can be measured, in this case the perturbation of neurite outgrowth. Positive controls and negative controls should always be included in such assays.
TempO-Seq (Templated Oligo-Sequencing) is based on hybridisation and sequencing technology. RNA-preserved from cells that have been exposed to the chemicals can undergo targeted sequencing. In this case, a ~3500 gene panel, established in close collaboration with the US National Toxicology program, was used. These assays were sensitive and able to cluster the groups of chemicals efficiently. Compounds could then be sorted into order by toxicity (i.e. calculating BMC values for the most relevant genes).
Conclusions
EU-ToxRisk is one of the projects driving the paradigm shift in toxicology towards an animal-free and mechanistic-based, integrated, and pragmatic approach to chemical safety assessment. The collaboration with ECHA also advances the understanding of both regulatory boundaries and science.
Highly toxic chemicals show more profound effects in reporter assays and their effects can be dependent on the cell type used. Phenotypic assays (e.g. neurite outgrowth assay) can be more sensitive than cytotoxicity. Finally, NAMs can contribute to the prioritisation of highly toxic chemicals.
The used assays were helpful to classify the chemicals based on different potency. Transcriptomics can reveal more about processes in the biological system than individual assays. This work is still ongoing and requires integration.
Questions, comments, and discussion
Marianne van der Hagen was wondering how many people were involved in the project? It would be 4 years of work for one person, however here there were many people involved, with different expertise.
Tomasz Sobański contrasted this case study and the approaches presented by Dr van de Water with those early presented by Dr Escher. He stressed that Dr Escher presented a concrete case study, specific hypothesis, proposed MoA, using very concrete assays, which are able to predict adverse outcomes to substantiate (agree/reject) these hypotheses, while Dr van de Water had a completely unknown MoA, unbiased chemicals where the main driver to choose these chemicals was availability of any reference data for repeated dose toxicity, and information about the industrial chemicals at the proper tonnage volume. He pointed out that this is purely an exploratory approach and that the project is trying to find out what kind of biology we need to cover to be successful. Later, this can be simplified. He also pointed out that assays do correlate, and even if they are sometimes not as perfect, their performance would resemble what Dr Paul Friedman referred to as a “gold standard” owing to the general variability of animal assays.
Owing to decreasing sequencing costs Marianne van der Hagen wondered if it is possible to run all chemicals with high-throughput transcriptomics. Dr van de Water commented that this is still expensive and requires quite a lot of samples per compound due to both the dose range and the models. The strategy proposed is to remove the most harmful chemicals using cheaper screening methods before entering transcriptomics. In the end, the most effective system would be for companies to deliver this data, supported by regulators to join forces (with part of the costs supplied by government incentives to speed up the process for the most harmful chemicals). This is already happening in the US (Tox21)
Dr Dirven stressed that repeated dose toxicity is based on many end points that regulatory bodies need to rely on. He wondered which of the points these approaches will not work for? What are the limitations of these NAMs approaches? There is obviously a need for further discussion about individual points, but Dr van de Water responded to this by asking how relevant for a human setting all these points are and if we really need to cover all those (i.e. 90-day studies). In addition, the goal with NAMs is to cover all biology programmes (many cell types with various gene expressions). There will always be a degree of uncertainty, but uncertainty is also present in the in vivo experiments, concluded Dr van de Water.
There was also discussion about the relevance of gene expression. At the end Dr van de Water expects only a handful of genes that will be relevant, with a link between gene expression and pathology, and that there is probably no need to show all the gene expression patterns.
Dr Sobański referred to a system that is currently being optimised but still performs pretty well at indicating which chemicals are likely to be carcinogenic, mutagenic and reprotoxic. We probably will be able to use such systems. He was not optimistic about the feasibility of replacing the repeated prenatal extended one generation tests, however.
Dr Lindeman agrees with Dr Sobański that there seems to be quite different tasks to design PODs that can be used to protect biology based on NAMs. At the end it seems one can select a combination of systems.
5.16. Orientation on relevant projects
RiskHUNT3R
By Prof Bob van de Water
Prof van de Water introduced a new project. RISK-HUNT3R - “RISK assessment of chemicals integrating HUman centric Next generation Testing strategies promoting the 3Rs”, is a European Research & Innovation project funded under Horizon 2020 programme. This 5-year project runs from June 2021 (with a budget of €22.9M, 37 partners involved across different sectors). RISK-HUNT3R builds on the outcomes and achievements of the EU-ToxRisk.
Throughout several case studies directed towards different modules, the project will develop, validate and implement integrated approaches to lead the way towards NGRA. Mechanism-based NAMs will be used in vitro and in silico systems with the endpoint relevant exclusively for human health. It will also assess uncertainty of these models. Developed in EU-ToxRisk NAMs toolbox will be fine-tuned and reused. The project will further optimise a strategy to assess chemical exposure, toxicokinetics, and toxicodynamics. It will extensively use ADME concepts (going from external to internal exposure, understanding the distribution and relevance), metabolic retrofitting, and high-throughput systems.
It will provide a framework for human relevant NGRA, based only on non-animal approaches to meet demands of future risk assessment. It will interact with chemical safety regulators to ensure future implementation. One of the goals is to develop IATAs that can be used for NGR testing, which will be commercially available at the end of the project.
OECD reporting - Omics technologies in chemical testing – OECD
By Prof John K. Colbourne
Prof Colbourne introduced ongoing active projects relating to the development of NAMs in the context of regulation. Prof Viant has been heavily involved in the OECD, and introduced us to several OECD projects, and programmes, especially in regard to the reporting of omics in risk assessment. Prof Colbourne also drew our attention to the OECD website (https://www.oecd.org/chemicalsafety/testing/omics.htm). This page is specifically devoted to explaining omics technologies in chemical testing, under the umbrella of chemical safety and biosafety. It presents an important overview of the different utilities of omics for the determination of chemical hazards and chemical risks. It also links to various other pages that are of relevance to both NAMs and to the application of omics with an explanation of why the OECD is working on omics. It also links to the IATAs, and is tied to the OECD hazard assessment programme, current and planned OECD activities and finally to an explanation of how people can engage with these OECD activities.
Accelerating the Pace of Chemical Risk Assessment (APCRA)
By Dr Tomasz Sobański
APCRA is an international government initiative which aims to collaborate and maintain dialogue on the scientific and regulatory needs for the application and acceptance of NAMs in regulatory decision making. Participants include members from North America, Europe, Asia and Australia’s governmental entities, as well as from OECD countries. Efforts have so far led to initiation of multiple collaborative case studies on the use of alternative methods in regulatory contexts.
APCRA holds regular workshops to discuss the development and application of NAMs for chemical risk assessment with international regulators. Case studies are initiated on an ongoing basis and continue to be discussed at annual workshops. The meetings have been for sharing data, knowledge, experience, and expertise among international government entities. This is also an arena for discussion on using NAMs in regulatory decisions including priority setting, hazard identification and risk assessment. It is also a platform for innovation and idea exchange between regulatory scientists, to discuss progress and barriers in applying new tools to prioritisation, as well as screening and quantitative risk assessments of different levels of complexity.
PrecisionTox
By Prof John K. Colbourne
The PrecisionTox project is funded by EU Horizon 2020 and will run for 5 years (started February 2021). It is led by the University of Birmingham. There are 15 different academic organisations and public research institutions involved. The mission of PrecisionTox is to draw causal links between molecular perturbations of adversity and to categorise variations in toxicological effects seen among different species and populations. This is in order to get a better understanding of the limits of exposure that lead to toxicological harm across biological systems. The project is focused on the identification and utility of these molecular key events that would be useful in a NAM context. It is tied into the AOP framework. The project aims to reconnect risk assessment for the protection of the environment with risk assessment for humans.
The goal of PrecisionTox is to improve chemical safety assessment to better protect human health and the environment by using non-traditional test species, multiple fields of knowledge, and powerful computational approaches to understand which chemicals are toxic and why. This goal is supported by three core concepts:
- Phylotoxicology: Replace traditional animal testing with an Evolutionarily Diverse Model Suite of organisms from multiple branches of the tree of life.
- Variation in susceptibility: determine safe levels of exposure to chemicals based on genetic variation.
- Embedded translation: Collaborate with regulators and other key stakeholders in project planning, selection of chemicals for investigation, and case studies for applying Precision Toxicology in policy and law.
PrecisionTox uses high-throughput testing methods across the evolutionarily diverse models of five biomedical model organisms and human cell lines to observe toxic responses (comparative toxicology). In this consortium, they are testing 250 chemicals systematically across different model species (i.e. C. elegans, Drosophila, Daphnia, embryos of Zebrafish and Xenopus), including human cell lines. It applies metabolomic and transcriptomic approaches to comparative toxicology samples to trace adverse outcomes via the molecular key events preceding them. It will assess a variation in susceptibility, by applying quantitative genetics and gene expression profiling to understand the variation in individual susceptibility and develop empirical exposure thresholds. Finally, it will use machine learning to identify biomarkers for molecular key events and create the dissemination and translation products for their use in regulatory analysis and application. This will be done in partnership with already existing risk managers and regulatory agencies. The project will provide new biomarker discoveries, PrecisionTox Data Commons (FAIR principle), and a NAM Toolbox.
The development of the project and outreach activities can be found on the PrecisionTox page precisiontox.org/.
PARC - The Partnership for the Assessment of Risk from Chemicals
By Dr Hubert Dirven
The Partnership for the Assessment of Risk from Chemicals (PARC) proposal was submitted to the European Commission for evaluation (and awarded in January 2022). PARC is a public partnership under EU Horizon Europe with a duration of 7 years. A 400 million Euro budget is allocated with approximately 50% -financing from the EU and 50% financing by the participating partners. 200 institutes from 28 countries are partners in PARC. The planned start of PARC is May 2022. Both human toxicology and ecotoxicology are focus areas in PARC.
PARC is an initiative where the European Union, with early involvement of Member States and Associated Countries, together with public partners (EU and National Risk Agencies, Universities, Public Research Organisations, etc.), commit to jointly support the development and implementation of a programme of research activities in relation to the assessment of the risk of chemicals. The European Green Deal and the Chemical Strategy for Sustainability are two of the central concepts in the development of this partnership. Regulatory needs from international and national regulatory authorities are the driving force behind PARC.
The aim of this Partnership is to enable all the scientific communities involved in chemical risk assessment, as well as risk managers and stakeholders, to have access to data according to the FAIR principle (Findable, Accessible, Interoperable, and Reusable). PARC will also make available concepts and toolboxes in the form of integration models. It will provide tools for the assessment of safe and sustainable by design substances. These tools will also facilitate early warning concepts (WP8).
In Norway, there are 8 institutes that are partners in PARC. NIPH has a coordinating role in Norway as the grant signatory and national hub coordinator. NIPH focuses mainly on WP4 (biomonitoring), WP5 (toxicology) and WP6 (regulatory implementation).
The WP4 aims at monitoring and measuring exposure both in humans and in the environment, considering the different sources, chemical fates and exposure pathways. It works on innovative analytical methods and tools, necessary for these types of study (i.e. surveys, measurement campaigns, sampling strategy).
The WP5 aims to contribute to increase the knowledgebase on hazards and to the development of hazard assessment methods for substances alone or in mixtures through the use of in vivo, in vitro, and in silico studies. It will contribute to the increased use of NAMs and provide data to fill gaps in our knowledge on natural toxins, legacy chemicals or of newly emerging hazards.
WP6 is focussed on innovation in regulatory risk assessment by contributing to the development of integrated testing and assessment approaches through crosscutting work with WP5 and WP4. In order to assess the transition to regulatory science, a review of existing regulatory assessment systems, their similarities and differences, will be carried out. This will help evaluate the strategies for transposing the new approaches, both in exposure science and hazard identification and characterisation, and will contribute to the future policies to apply the Chemical Strategy.
PARC will support the development of laboratory capacities and interinstitutional networking in the different fields of activities by identifying existing and yet-to-be-developed networks, supporting the implementation of standardisation approaches, and evaluating the reproducibility of performances and their monitoring (WP9). This would result in developing IATAs.
ONTOX
By Dr Hubert Dirven
The ONTOX project started in May 2021 and will last until April 2026. It is funded under the EU Horizon 2020 framework and has an overall budget of €17M. The project is coordinated by Prof Mathieu Vinken from the Vrije Universiteit Brussel in Belgium.
The vision of the ONTOX is to advance human risk assessment of chemicals without the use of animals, in line with the principles of 21st century toxicity testing and NGRA (3Rs and NAMs). ONTOX's goal is to deliver a proof-of-concept strategy to create NAMs to predict systemic repeated dose toxicity effects that (upon combination with tailored exposure assessment) will enable human risk assessment. For this purpose, ONTOX focuses on 6 specific NAMs addressing adversities in the liver (steatosis and cholestasis), kidneys (tubular necrosis and crystalopathy) and developing brain (neural tube closure and cognitive function defects) induced by a variety of chemicals.
The NAMs will each consist of a computational system based on artificial intelligence (AI) and will be primarily fed by available biological, mechanistic, toxicological, epidemiological, physico-chemical and kinetic data. Data will be consecutively integrated in physiological maps, quantitative AOP networks and ontology frameworks. Data gaps will be filled by targeted in vitro and in silico testing.
Development of the project and outreach activities can be found on the ONTOX page: ontox-project.eu/.
Research cluster ASPIS
By Prof John K. Colbourne, Prof Bob van de Water and by Dr Hubert Dirven
ASPIS cluster is a joint collaboration of the H2020 funded projects ONTOX, PrecisionTox, RISK-HUNT3R and represents Europe’s €60M effort towards the sustainable, animal-free and reliable chemical risk assessment of tomorrow. ASPIS stands for (“Animal-free Safety assessment of chemicals: Project cluster for Implementation of novel Strategies”).
5.17. H2020 PrecisionTox omics-based G/RAx case study in the making
By Prof Mark Viant
In his talk, Prof Viant revisited the PrecisionTox project, and work package six in particular, which focuses on regulatory analysis and application.
Two of the main objectives in PrecisionTox work package six are 1) to develop specific case studies that demonstrate the use of omics/molecular biomarkers in grouping and read-across, and 2) produce draft guidance for the use of omics/molecular biomarkers in grouping and read-across, describing “acceptable practice” and how to report and interpret the findings, and finally to maximise the transparency and ease of use by all stakeholders.
The first objective highlights the importance of using case studies for developing NAMs, especially for translating and evaluating NAMs for regulatory purposes. The second objective emphasises that we need to start talking about what is acceptable practice in terms of NAMs. We should decide on how to report and how to interpret the findings from NAMs to obtain consistency between laboratories. Ensuring transparency in these approaches is a critical step towards their usability by both regulators and industry.
Before the workshop, the participants were asked to provide examples of where they are currently experiencing challenges in grouping /read-across, in particular when evidence to support the grouping hypothesis is weak or not consistent. These examples were considered by the workshop trainers in an attempt to identify which could be proposed as PrecisionTox case studies. The Swedish Chemical Agency (Ilona Silins) suggested a case study on acrylamides, and the Norwegian Institute of Public Health (Birgitte Lindeman) suggested a case study on imidazoles. These real-world case studies are now under consideration to be included in PrecisionTox as omics-based grouping and read-across case studies.
Finally, and most importantly, Prof Viant invited the workshop participants to collaborate on the grouping and read-across case study(ies) with Michabo Health Science Ltd, that he represents to:
- Learn more about NAMs from a team of experts
- Contribute to a case study to maximise its likelihood of delivering value to the regulatory community
- Advise on the draft guidance document to ensure it covers the topics of relevance to regulators
- Join a future workshop(s) where the results are presented and discussed
6. Acknowledgements
We would like to thank David Epps (University of Birmingham) and Elena Sostare (Michabo Health Science Ltd) who contributed to the pre-workshop survey and slides to present the survey results, and Dr Camilla Svendsen (Norwegian Institute of Public Health) for proofreading the report.
7. Conflict of interest
Professors Mark Viant and John K. Colbourne are employees of the University of Birmingham and Founders and Directors of Michabo Health Science (MHS) Ltd., a spin-out company of the University of Birmingham. MHS also operates as a trading division of University of Birmingham Enterprise Ltd., a wholly owned subsidiary of the University of Birmingham. MHS provides scientific consultancy services in NAMs specialising in omics technologies, computational toxicology, and training.
A framework agreement exists between ECHA and Michabo Health Science Ltd. with the purpose of NAM competence development. NORAP (NOrdic Risk Assessment Project) and NKIG (Nordic classification group) were involved in the planning of this workshop and have agreed to contribute to the Nordic workshop on NAMs.
Rapporteurs
Dr Marcin W. Wojewodzic, Norwegian Institute of Public Health, Marcin.Wojewodzic@fhi.no
Dr Monica Andreassen, Norwegian Institute of Public Health, Monica.Andreassen@fhi.no
Appendix 1: Programme
Day 1: Tuesday November 9
| Time | Mins | Title | Description | Presenter |
| 09:00 | 10’ | Introduction & Objectives |
| Marianne van der Hagen (Norwegian Environment Agency) |
| 09:10 | 20’ | Poll 1 | Current level of knowledge about New Approach Methodologies (NAMs)
| Hubert Dirven (Norwegian Institute of Public Health) |
| 09:30 | 30’ | Introduction to NAMs |
| John Colbourne & Mark Viant (Michabo Health Science Ltd, UK) |
| 10:00 | 30’ | Introduction to omics |
| Mark Viant & John Colbourne (Michabo Health Science Ltd, UK) |
| 10:30 | 10’ | BREAK | ||
| 10:40 | 30’ | Towards regulatory applications of molecular mechanistic data I |
| Mark Viant (Michabo Health Science Ltd, UK) |
| 11:10 | 30’ | Towards regulatory applications of molecular mechanistic data II |
| John Colbourne (Michabo Health Science Ltd, UK) |
| 11:40 | 20’ | BREAK | ||
| 12:00 | 60’ | Invited talk - Introducing the CompTox Chemicals Dashboard |
| Grace Patlewicz & Antony Williams (U.S. Environmental Protection Agency) |
| 13:00 | 30’ | Questions & Answers | Marianne van der Hagen (Norwegian Environment Agency) |
Day 2: Wednesday November 10
| Time (CET) | Mins | Title | Description | Presenter |
| 09:00 | 15’ | Introduction & Highlights from Day 1 | Birgitte Lindeman (Norwegian Institute of Public Health) | |
| 09:15 | 20’ | Challenges of NAMs in the Real World 1 |
| Hubert Dirven (Norwegian Institute of Public Health) |
| 09:35 | 20’ | Challenges of NAMs in the Real World 2 |
| Magda Sachana (OECD) |
| 09:55 | 35’ | Group Discussion |
| Hubert Dirven (Norwegian Institute of Public Health) |
| 10:30 | 10’ | BREAK | ||
| 10:40 | 60’ | Experience of ECHA in applying NAMs in a regulatory context |
| Tomasz Sobański (ECHA) Mark Viant (Michabo Health Science Ltd, UK) Mounir Bouhifd (ECHA) |
| 11:40 | 35’ | BREAK | ||
| 12:15 | 45’ | Invited talk - Quantitative and qualitative reproducibility of study-level and organ-level effects in repeat dose animal studies |
| Katie Paul Friedman (U.S. Environmental Protection Agency) |
| 13:00 | 30’ | Questions & Answers | Marianne van der Hagen (Norwegian Environment Agency) | |
| 13:30 |
Day 3: Thursday November 11
| Time (CET) | Mins | Title | Description | Presenter |
| 09:00 | 15’ | Introduction & Highlights from Day 2 | Birgitte Lindeman (Norwegian Institute of Public Health) | |
| 09:15 | 45’ | Grouping and Read-Across Case study from EUToxRISK |
| Sylvia Escher (Fraunhofer Institute for Toxicology and Experimental Medicine) |
| 10:00 | 30’ | ECHA’s experiences with applying NAMs in the assessments of chemical groups |
| Tomasz Sobański (ECHA) |
| 10:30 | 10’ | BREAK | ||
| 10:40 | 60’ | Invited talk - EUToxRisk Case Study 11 |
| Bob van de Water (Leiden University) |
| 11:40 | 20’ | BREAK | ||
| 12:00 | 30’ | Orientation on Relevant Projects |
| John Colbourne (University of Birmingham), Tomasz Sobański (ECHA), Hubert Dirven (Norwegian Institute of Public Health), Bob van de Water (Leiden University) |
| 12:30 | 10’ | H2020 PrecisionTox omics-based G/RAx case study in the making |
| Mark Viant (Michabo Health Science Ltd) |
| 12:40 | 20’ | Poll 2 | As the workshop draws to a close, has the workshop successfully provided added knowledge about New Approach Methodologies (NAMs)? | Hubert Dirven (Norwegian Institute of Public Health) |
| 13:00 | 30’ | Wrap up |
| Marianne van der Hagen (Norwegian Environment Agency) Daniel Borg (Swedish Chemicals Agency) |
| 13:30 |
Appendix 2: Supplementary figures
Results from the pre-workshop poll

Figure 1. Which New approach methods have you 1) heard about or 2) have some experiences with?

Figure 2. Where do you see the largest opportunities of NAMs in the next 5 years? Score from 1 to 5 (1= no opportunity, 5 = large opportunity).

Figure 3. What is the biggest hurdle using NAMs in your hazard assessments? Score from 1 – 5 (1 = no hurdle, 5 = a big hurdle).

Figure 4. Results from the poll questions: A) Are you informed about freely available data produced by New Approach Methodologies (NAMs)? B) Which data sources are you aware of?
Results from the post-workshop poll
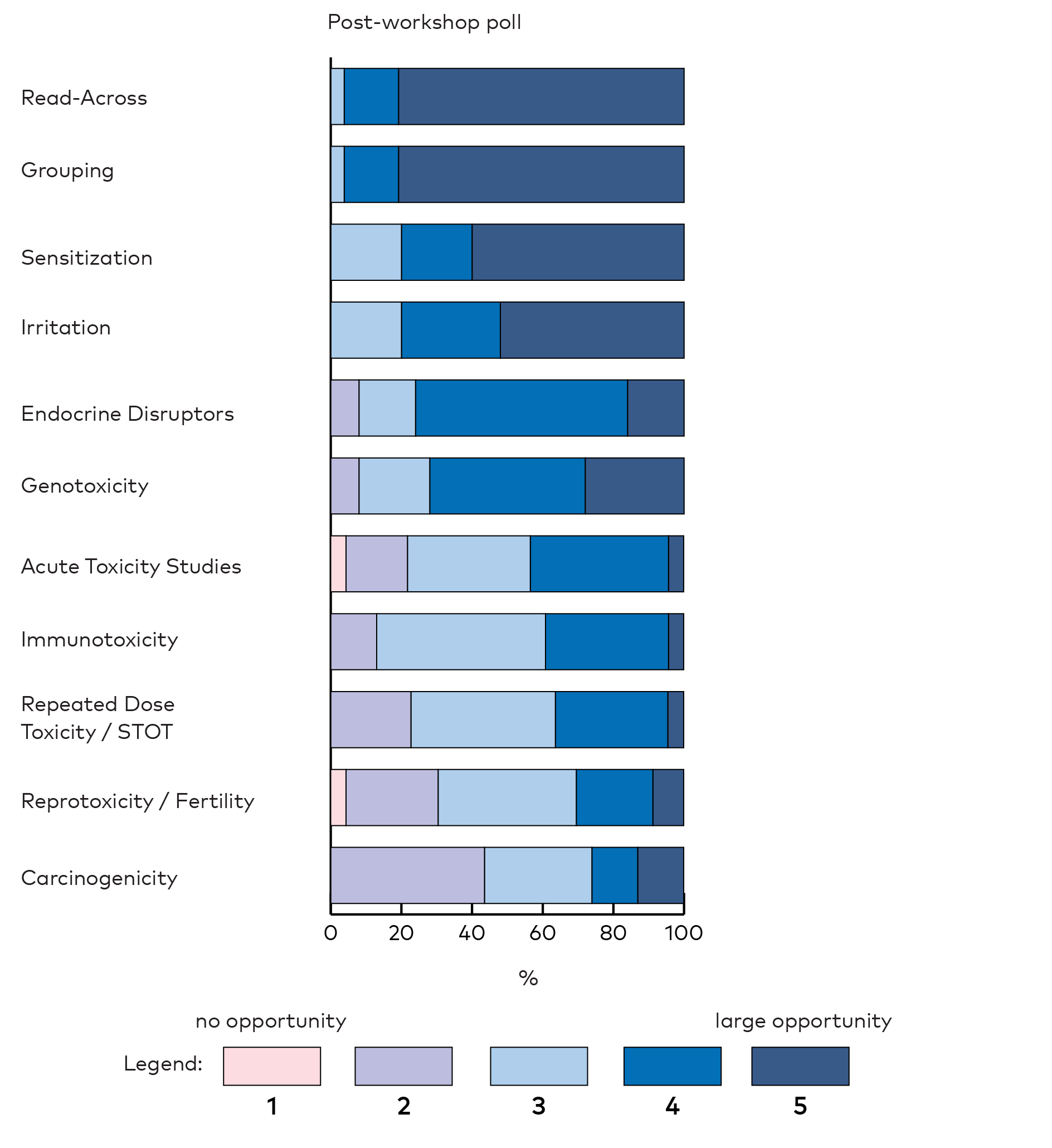
Figure 5. Where do you see the largest opportunities of NAMs in the next 5 years? Score from 1 to 5 (1= no opportunity, 5 = large opportunity).
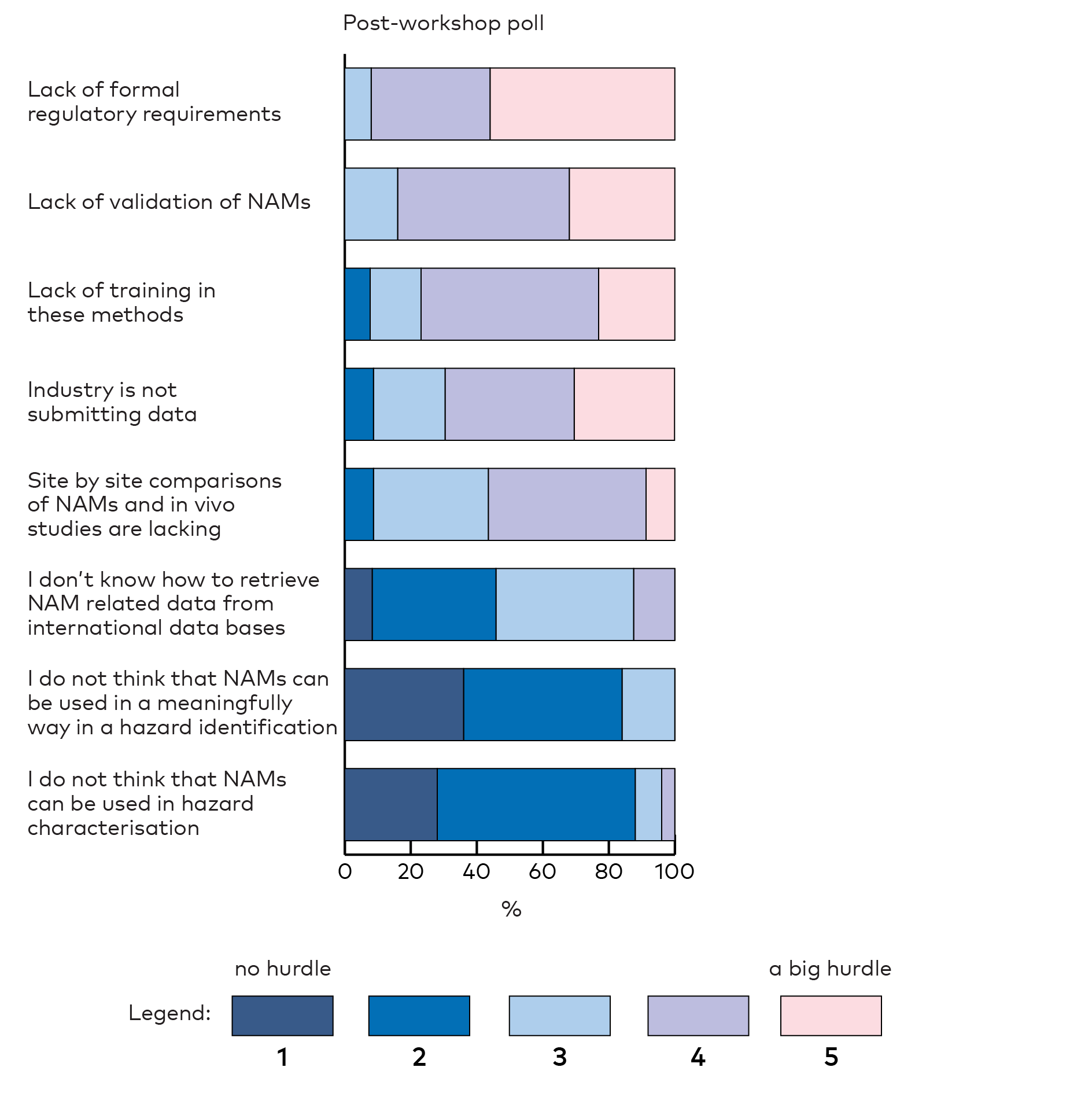
Figure 6. What is the biggest hurdle using NAMs in your hazard assessments? Score from 1 – 5 (1 = no hurdle, 5 = a big hurdle).
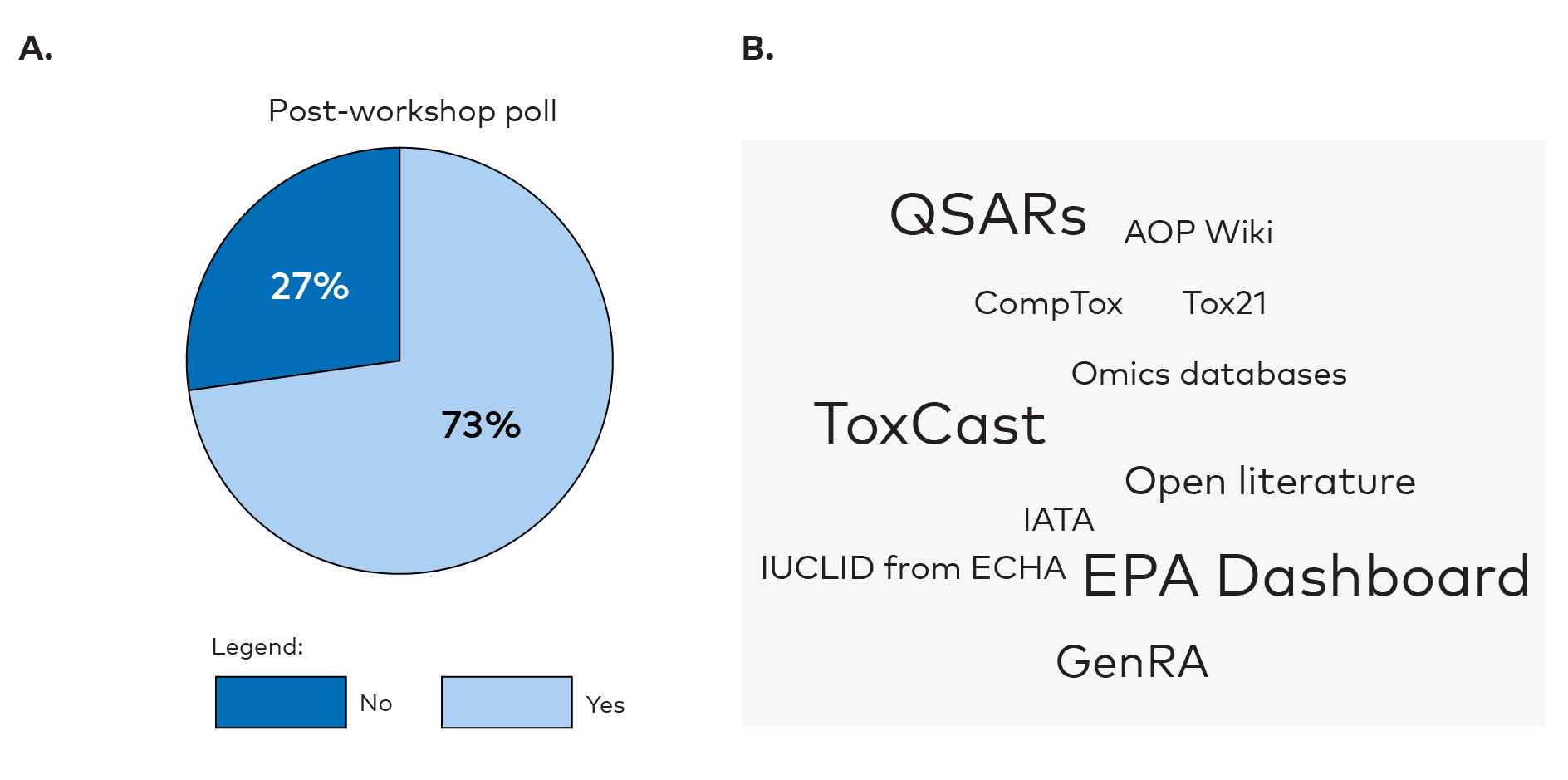
Figure 7. Results from the post-workshop poll questions: A) Are you informed about freely available data produced by New Approach Methodologies (NAMs)? B) Which data sources are you aware of?
About this publication
Nordic Workshop on New Approach Methodologies (NAMs)
for Grouping and Read-Across under REACH and CLP
Dr Marcin W. Wojewodzic
Dr Monica Andreassen
ISBN 978-92-893-7313-5 (PDF)
ISBN 978-92-893-7314-2 (ONLINE)
http://dx.doi.org/10.6027/temanord2022-526
TemaNord 2022:526
ISSN 0908-6692
© Nordic Council of Ministers 2022
Illustration: Ella Maru Studio
Published: 20/5/2022
Disclaimer
This publication was funded by the Nordic Council of Ministers. However, the content does not necessarily reflect the Nordic Council of Ministers’ views, opinions, attitudes or recommendations.
Rights and permissions
This work is made available under the Creative Commons Attribution 4.0 International license (CC BY 4.0) https://creativecommons.org/licenses/by/4.0.
Translations: If you translate this work, please include the following disclaimer: This translation was not produced by the Nordic Council of Ministers and should not be construed as official. The Nordic Council of Ministers cannot be held responsible for the translation or any errors in it.
Adaptations: If you adapt this work, please include the following disclaimer along with the attribution: This is an adaptation of an original work by the Nordic Council of Ministers. Responsibility for the views and opinions expressed in the adaptation rests solely with its author(s). The views and opinions in this adaptation have not been approved by the Nordic Council of Ministers.
Third-party content: The Nordic Council of Ministers does not necessarily own every single part of this work. The Nordic Council of Ministers cannot, therefore, guarantee that the reuse of third-party content does not infringe the copyright of the third party. If you wish to reuse any third-party content, you bear the risks associated with any such rights violations. You are responsible for determining whether there is a need to obtain permission for the use of third-party content, and if so, for obtaining the relevant permission from the copyright holder. Examples of third-party content may include, but are not limited to, tables, figures or images.
Photo rights (further permission required for reuse):
Any queries regarding rights and licences should be addressed to:
Nordic Council of Ministers/Publication Unit
Ved Stranden 18
DK-1061 Copenhagen
Denmark
pub@norden.org
Nordic co-operation
Nordic co-operation is one of the world’s most extensive forms of regional collaboration, involving Denmark, Finland, Iceland, Norway, Sweden, and the Faroe Islands, Greenland and Åland.
Nordic co-operation has firm traditions in politics, economics and culture and plays an important role in European and international forums. The Nordic community strives for a strong Nordic Region in a strong Europe.
Nordic co-operation promotes regional interests and values in a global world. The values shared by the Nordic countries help make the region one of the most innovative and competitive in the world.
The Nordic Council of Ministers
Nordens Hus
Ved Stranden 18
DK-1061 Copenhagen
pub@norden.org
Read more Nordic publications on www.norden.org/publications